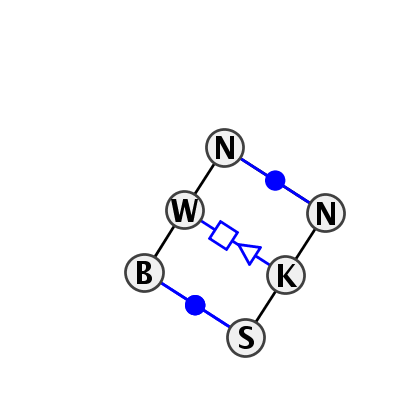
Motif IL_01903.1 Version IL_01903.1 of this group appears in releases 2.0 to 2.0
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name | 1 | 2 | 3 | break | 4 | 5 | 6 | 1-5 | 1-6 | 2-5 | 3-4 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | ➜ | 5AJ0 | 0.5344 | 1 | A2 | LSU rRNA | C | 2625 | A | 2626 | G | 2627 | * | C | 2664 | G | 2665 | G | 2667 | cWW | cWW | |||
| 2 | ➜ | 3JCS | 0.2947 | 2 | 2 | LSU rRNA | C | 1094 | A | 1095 | G | 1096 | * | C | 1114 | G | 1115 | G | 1118 | cWW | tHS | cWW | ||
| 3 | ➜ | 5FDU | 0.3024 | 2 | 1A | LSU rRNA | C | 2309 | A | 2310 | G | 2311 | * | C | 2329 | G | 2330 | G | 2333 | cWW | tHS | cWW | ||
| 4 | ➜ | 4V9F | 0.2896 | 2 | Isolated tHS basepair with bulges | 0 | LSU rRNA | C | 2331 | A | 2332 | G | 2333 | * | C | 2351 | G | 2352 | G | 2355 | cWW | tHS | cWW | |
| 5 | ➜ | 4V8P | 0.2478 | 2 | F1 | LSU rRNA | C | 2655 | A | 2656 | G | 2657 | * | C | 2675 | G | 2676 | G | 2679 | cWW | tHS | cWW | ||
| 6 | ➜ | 5AN9 | 0.2655 | 2 | N | LSU rRNA | C | 2999 | A | 3000 | G | 3001 | * | C | 3019 | G | 3020 | G | 3023 | cWW | tHS | cWW | ||
| 7 | ➜ | 5AJ0 | 0.2135 | 2 | A2 | LSU rRNA | C | 4205 | A | 4206 | G | 4207 | * | C | 4225 | G | 4226 | G | 4229 | cWW | tHS | cWW | ||
| 8 | ➜ | 3J79 | 0.2113 | 2 | A | LSU rRNA | C | 3005 | A | 3006 | A | 3007 | * | U | 3025 | G | 3026 | G | 3029 | cWW | tHS | cWW | ||
| 9 | ➜ | 4IOA | 0.2612 | 2 | Isolated tHS basepair with bulges | X | LSU rRNA | C | 2276 | A | 2277 | A | 2278 | * | U | 2296 | G | 2297 | G | 2300 | cWW | tHS | cWW | |
| 10 | ➜ | 4V88 | 0.2520 | 2 | A5 | LSU rRNA | C | 2666 | A | 2667 | U | 2668 | * | A | 2686 | G | 2687 | G | 2690 | cWW | tHS | cWW | ||
| 11 | ➜ | 4V91 | 0.2704 | 2 | 1 | LSU rRNA | C | 2666 | A | 2667 | U | 2668 | * | A | 2686 | G | 2687 | G | 2690 | cWW | tHS | cWW | ||
| 12 | ➜ | 3J7P | 0.2144 | 2 | 5 | LSU rRNA | C | 4243 | A | 4244 | G | 4245 | * | C | 4263 | G | 4264 | G | 4267 | cWW | tHS | cWW | ||
| 13 | ➜ | 5FDU | 0.0000 | 1 | 1A | LSU rRNA | C | 2103 | A | 2104 | G | 2105 | * | C | 2248 | G | 2249 | G | 2251 | cWW | tHS | cWW | ||
| 14 | ➜ | 4V9F | 0.1088 | 1 | Isolated tHS basepair with bulges | 0 | LSU rRNA | C | 2122 | A | 2123 | G | 2124 | * | C | 2269 | G | 2270 | G | 2272 | cWW | tHS | cWW | |
| 15 | ➜ | 3J9W | 0.1326 | 1 | AA | SSU rRNA | C | 1469 | A | 1470 | G | 1471 | * | U | 1449 | G | 1450 | G | 1452 | cWW | tHS | cWW | ||
| 16 | ➜ | 3JCS | 0.1797 | 1 | 2 | LSU rRNA | C | 669 | A | 670 | G | 671 | * | U | 1032 | G | 1033 | G | 1035 | cWW | tHS | cWW | ||
| 17 | ➜ | 5FDU | 0.1790 | 1 | 1A | LSU rRNA | C | 312 | A | 313 | G | 314 | * | U | 374 | G | 375 | G | 377 | cWW | tHS | cWW | ||
| 18 | ➜ | 3J7P | 0.2054 | 1 | 5 | LSU rRNA | U | 3927 | A | 3928 | G | 3929 | * | U | 4181 | G | 4182 | G | 4184 | cWW | tHS | cWW | ||
| 19 | ➜ | 4IOA | 0.2998 | 1 | Isolated tHS basepair with bulges | X | LSU rRNA | U | 2064 | A | 2065 | G | 2066 | * | C | 2215 | G | 2216 | G | 2218 | cWW | tHS | cWW | |
| 20 | ➜ | 5J7L | 0.2787 | 1 | Isolated tHS basepair with bulges | DA | LSU rRNA | U | 2081 | A | 2082 | G | 2083 | * | U | 2236 | G | 2237 | G | 2239 | cWW | tHS | cWW | |
| 21 | ➜ | 3J79 | 0.2984 | 1 | A | LSU rRNA | U | 2716 | A | 2717 | G | 2718 | * | U | 2943 | G | 2944 | G | 2946 | cWW | tHS | cWW | ||
| 22 | ➜ | 5AJ0 | 0.2770 | 1 | A2 | LSU rRNA | U | 3898 | A | 3899 | G | 3900 | * | U | 4143 | G | 4144 | G | 4146 | cWW | tHS | cWW | ||
| 23 | ➜ | 4V91 | 0.3113 | 1 | 1 | LSU rRNA | U | 2423 | A | 2424 | G | 2425 | * | U | 2604 | G | 2605 | G | 2607 | cWW | tHS | cWW | ||
| 24 | ➜ | 5AN9 | 0.3254 | 1 | N | LSU rRNA | U | 2689 | A | 2690 | G | 2691 | * | U | 2937 | G | 2938 | G | 2940 | cWW | tHS | cWW | ||
| 25 | ➜ | 4V88 | 0.2930 | 1 | A5 | LSU rRNA | U | 2423 | A | 2424 | G | 2425 | * | U | 2604 | G | 2605 | G | 2607 | cWW | tHS | cWW | ||
| 26 | ➜ | 5A2Q | 0.6143 | 0 | 3 | IRES | G | 222 | A | 223 | G | 224 | * | U | 175 | G | 176 | C | 177 | cWW | tHS | cWW | ||
| 27 | ➜ | 4KYY | 0.7560 | 0 | B+A | RNA 17-mer | U | 8 | U | 9 | G | 10 | * | U | 8 | U | 9 | G | 10 | cWW | ncWW | cWW | ||
| 28 | ➜ | 4BTS | 0.8099 | 1 | DA | SSU rRNA | C | 492 | A | 494 | C | 495 | * | G | 477 | G | 478 | G | 479 | ncWW | cWW | tHS | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UAG*UGGG | 7 |
| CAG*CGUAG | 3 |
| CAU*AGUAG | 2 |
| CAA*UGUAG | 2 |
| CAG*UGGG | 2 |
| CAG*CGGG | 2 |
| CAG*CGUGG | 2 |
| GAG*UGC | 1 |
| UUG*UUG | 1 |
| CUAC*GGG | 1 |
| UAG*CGGG | 1 |
| CAG*CGGAG | 1 |
| CAG*CGUG | 1 |
| CAG*UGAG | 1 |
| CAG*CGAAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| A*GG | 12 |
| A*GUA | 7 |
| A*GUG | 2 |
| U*U | 1 |
| UA*G | 1 |
| A*G | 1 |
| A*GA | 1 |
| A*GU | 1 |
| A*GGA | 1 |
| A*GAA | 1 |
Release history
| Release | 2.0 |
|---|---|
| Date | 2017-04-24 |
| Status | New id, 2 parents |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- Isolated tHS basepair with bulges (5)
- Basepair signature
- cWW-tHS-cWW
- Heat map statistics
- Min 0.09 | Avg 0.39 | Max 0.99
Coloring options: