- Structure title
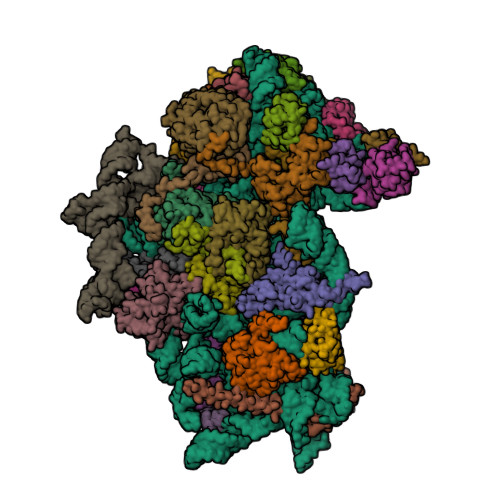
- Structure of the HCV IRES binding to the 40S ribosomal subunit, closed conformation. Structure 4(delta dII)
- Authors
- Brown, Z.P., Abaeva, I.S., De, S., Hellen, C.U.T., Pestova, T.V., Frank, J.
- Release date
- 2022-07-13
- Experimental technique
- ELECTRON MICROSCOPY
- Resolution
- 4.8 Å
- Chains
-
2 RNA chains from
Oryctolagus cuniculus, hepatitis C virus genotype 1a
-
34 other chains
- Nucleic acid compounds
18S rRNA, HCV IRES
- Other compounds
40S ribosomal protein SA, eS1, uS5 (S2), uS3, 40S ribosomal protein S4, uS7, eS6, 40S ribosomal protein S7, eS8, uS4, eS10, uS17, eS12, uS15, uS11, uS19, uS9, eS17, uS13, eS19, uS10, 40S ribosomal protein S21, uS8, uS12, 40S ribosomal protein S24, eS25 (S25), eS26 (S26), eS27, eS28, eS29, eS30, 40S ribosomal protein S27a, RACK1, 60s ribosomal protein l41
Pairwise Interactions
Interactions annotated by
FR3D:
View all
RNA 3D Motifs
In the current release of the
RNA 3D Motif Atlas, 7SYJ contains:
-
48 hairpin loops from 0 motif groups
-
107 internal loops from 0 motif groups
-
29 junction loops from 0 motif groups
More
3D Redundancy
7SYJ has chains which belong to the following equivalence classes in the current representative set:
Similar structures
None found