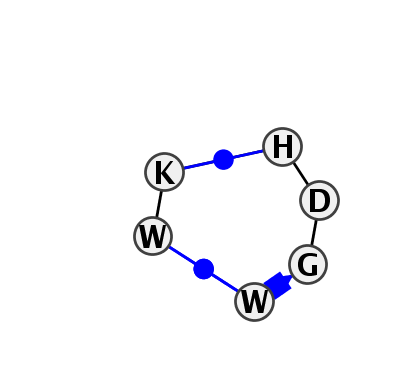
Motif IL_64441.1 Version IL_64441.1 of this group appears in releases 2.0 to 2.0
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | break | 3 | 4 | 5 | 6 | 1-6 | 2-3 | 5-6 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_4BTS_320 | 4BTS | 0.4998 | 2 | DA | SSU rRNA | U | 1229 | U | 1230 | * | A | 1197 | A | 1198 | G | 1201 | A | 1202 | cWW | cWW | ncSH |
| 2 | IL_4V9F_009 | 4V9F | 0.2682 | 2 | 0 | LSU rRNA | A | 248 | G | 249 | * | C | 260 | A | 261 | G | 264 | U | 265 | cWW | cWW | |
| 3 | IL_4V8P_306 | 4V8P | 0.0000 | 2 | F1 | LSU rRNA | A | 121 | U | 122 | * | A | 144 | A | 145 | G | 148 | U | 149 | cWW | cWW | |
| 4 | IL_4V88_248 | 4V88 | 0.1487 | 2 | A5 | LSU rRNA | A | 123 | U | 124 | * | A | 144 | G | 145 | G | 148 | U | 149 | cWW | cWW | |
| 5 | IL_4V91_007 | 4V91 | 0.2225 | 2 | 1 | LSU rRNA | A | 123 | U | 124 | * | A | 144 | G | 145 | G | 148 | U | 149 | cWW | cWW | |
| 6 | IL_5A2Q_022 | 5A2Q | 0.8087 | 2 | 2 | SSU rRNA | U | 393 | G | 394 | * | U | 366 | U | 367 | G | 370 | A | 371 | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AU*AGUUGU | 2 |
| UU*AAGGGA | 1 |
| AG*CAAUGU | 1 |
| AU*AAUUGU | 1 |
| UG*UUUCGA | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| *GUUG | 2 |
| *AGGG | 1 |
| *AAUG | 1 |
| *AUUG | 1 |
| *UUCG | 1 |
Release history
| Release | 2.0 |
|---|---|
| Date | 2017-04-24 |
| Status | New id, no parents |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (6)
- Basepair signature
- cWW-cSH-R-cWW
- Heat map statistics
- Min 0.15 | Avg 0.43 | Max 0.97
Coloring options: