- Structure title
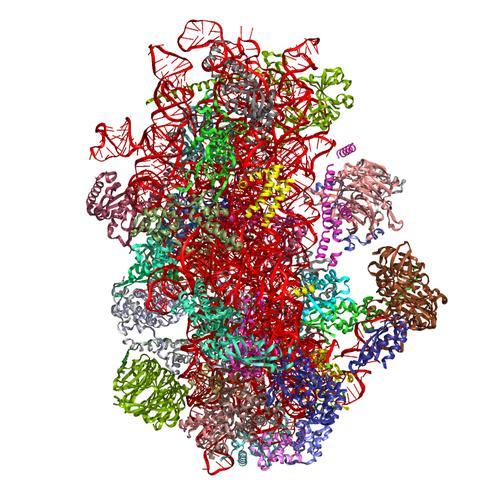
- Structure of a partial yeast 48S preinitiation complex in open conformation
- Authors
- Release date
- 2015-08-12
- Experimental technique
- ELECTRON MICROSCOPY
- Resolution
- 6.0 Å
- Chains
-
3 RNA chains from
Saccharomyces cerevisiae, Kluyveromyces lactis, synthetic construct
-
44 other chains
- Nucleic acid compounds
Met-tRNAi, 18S rRNA, mRNA
- Other compounds
uS2, eS1, uS5, uS3, eS4, uS7, eS6, eS7, eS8, uS4, eS10, uS17, eS12, uS15, uS11, uS19, uS9, eS17, uS13, eS19, uS10, eS21, uS8, uS12, eS24, eS25, eS26, eS27, eS28, uS14, eS30, eS31, RACK1, eL41, eIF1A, eIF2 alpha, eIF2 gamma, eIF2 beta, eIF1, eIF3a, eIF3c, eIF3i, eIF3b, eIF3g
Pairwise Interactions
Interactions annotated by
FR3D:
View all
RNA 3D Motifs
In the current release of the
RNA 3D Motif Atlas, 3jaq contains:
-
41 hairpin loops from 0 motif groups
-
93 internal loops from 0 motif groups
-
27 junction loops from 0 motif groups
More
3D Redundancy
This PDB is not included in any representative set equivalence class. Possibly, this PDB doesn't contain any complete nucleotides.
Similar structures
None found