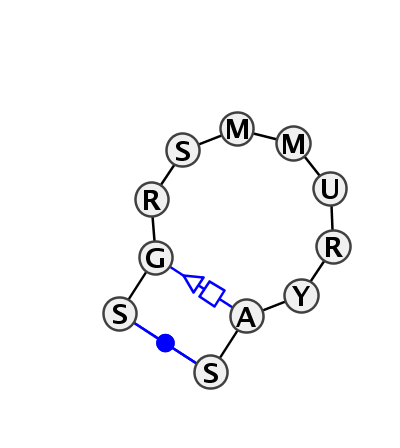
Motif HL_19162.1 Version HL_19162.1 of this group appears in releases 3.2 to 3.11
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 1-11 | 2-10 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_3IVK_001 | 3IVK | 0.1497 | 1 | M | class I ligase product | G | 1 | G | 2 | A | 3 | C | 5||A | A | 6 | C | 7 | U | 8 | A | 9 | U | 10 | A | 11 | C | 12 | cWW | tSH |
| 2 | HL_3HHN_006 | 3HHN | 0.0000 | 1 | E | Class I ligase ribozyme, self-ligation product | G | 1 | G | 2 | A | 3 | C | 5 | A | 6 | C | 7 | U | 8 | A | 9 | U | 10 | A | 11 | C | 12 | cWW | tSH |
| 3 | HL_3DHS_005 | 3DHS | 0.5586 | 1 | A | RNase P class B | C | 313 | G | 314 | G | 315 | G | 317 | C | 318 | A | 319 | U | 320 | G | 321 | C | 322 | A | 323 | G | 324 | cWW | ntSH |
| 4 | HL_2A64_006 | 2A64 | 0.8303 | 1 | A | RNase P class B | C | 313 | G | 314 | G | 315 | G | 317 | C | 318 | A | 319 | U | 320 | G | 321 | C | 322 | A | 323 | G | 324 | cWW | ntSH |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GGAACACUAUAC | 2 |
| CGGCGCAUGCAG | 2 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAACACUAUA | 2 |
| GGCGCAUGCA | 2 |
Release history
| Release | 3.3 | 3.4 | 3.5 | 3.6 | 3.7 | 3.8 | 3.9 | 3.10 | 3.2 | 3.11 |
|---|---|---|---|---|---|---|---|---|---|---|
| Date | 2018-03-08 | 2018-04-06 | 2018-05-04 | 2018-06-01 | 2018-06-29 | 2018-07-27 | 2018-08-24 | 2018-09-21 | 2018-09-26 | 2018-10-19 |
| Status | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | New id, no parents | Exact match |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (4)
- Basepair signature
- cWW-tSH-R-R-R-R-R-R-R
- Heat map statistics
- Min 0.15 | Avg 0.43 | Max 0.86
Coloring options: