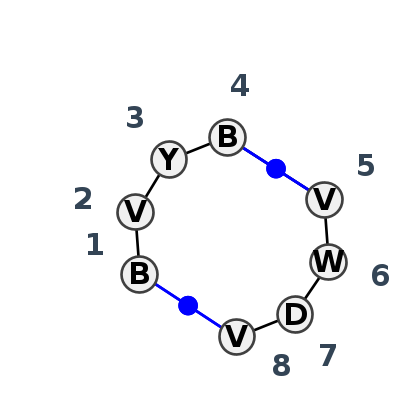
Motif IL_65740.2 Version IL_65740.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | break | 5 | 6 | 7 | 8 | 1-8 | 2-7 | 3-6 | 4-5 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_3NPQ_006 | 3NPQ | 0.7267 | 2 | A | SAH (S-adenosyl-L-homocysteine) riboswitch aptamer | C | 26 | C | 27 | C | 28 | G | 31 | * | C | 16 | A | 17 | A | 18 | G | 19 | cWW | ncWW | tSH | cWW | |
| 2 | IL_4X4T_003 | 4X4T | 0.0000 | 0 | G | G70A tRNA minihelix ending in CCACCA | G | 1||A | G | 2||A | C | 3||A | C | 4||A | * | G | 28||A | A | 29||A | 5BU | 30||A | C | 31||A | cWW | ncWWa | ncWW | cWW | |
| 3 | IL_8BH9_001 | 8BH9 | 0.1276 | 0 | B | RNA (12-mer) | C | 500 | A | 501 | U | 502 | U | 503 | * | A | 430 | A | 431 | U | 432 | G | 433 | cWW | cHW | cWW | ||
| 4 | IL_4WF9_110 | 4WF9 | 0.3299 | 0 | Tandem non-canonical cWW pairs | Y | 5S rRNA | U | 107 | G | 108 | C | 109 | C | 110 | * | G | 5 | U | 6 | G | 7 | A | 8 | cWW | ncWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CCCAGG*CAAG | 1 |
| GGCC*GA(5BU)C | 1 |
| CAUU*AAUG | 1 |
| UGCC*GUGA | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| CCAG*AA | 1 |
| GC*A(5BU) | 1 |
| AU*AU | 1 |
| GC*UG | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (3)
- Tandem non-canonical cWW pairs (1)
- Basepair signature
- cWW-L-R-L-R-cWW
- Heat map statistics
- Min 0.13 | Avg 0.39 | Max 0.87
Coloring options: