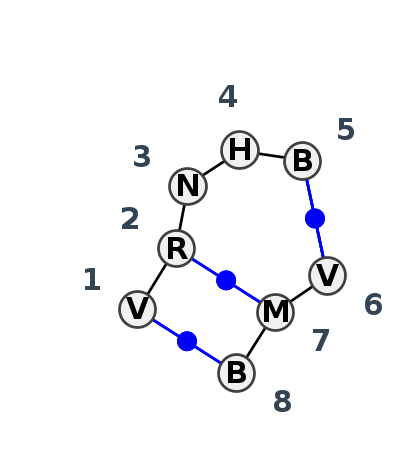
Motif IL_70784.2 Version IL_70784.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | break | 6 | 7 | 8 | 1-8 | 2-7 | 3-4 | 3-5 | 4-7 | 5-6 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_3OXE_002 | 3OXE | 0.9528 | 0 | A | Glycine riboswitch | C | 31 | G | 32 | A | 33 | A | 34 | G | 35 | * | C | 70 | A | 71 | G | 72 | cWW | tHS | cWW | ||||
| 2 | IL_9H3G_015 | 9H3G | 0.3214 | 0 | A | LSU rRNA | G | 234 | A | 235 | C | 236 | C | 237 | C | 238 | * | G | 174 | A | 175 | C | 176 | cWW | cwW | ncWWa | cWW | |||
| 3 | IL_6M0X_001 | 6M0X | 0.1593 | 0 | B | RNA (71-MER) | A | 38 | A | 39 | G | 40 | C | 41 | U | 42 | * | A | 29 | C | 30 | U | 31 | cWW | ncWWa | ncSH | ncwW | cWW | ||
| 4 | IL_7EL1_001 | 7EL1 | 0.0000 | 0 | B | RNA (73-MER) | A | 42 | A | 43 | U | 44 | C | 45 | U | 46 | * | A | 29 | C | 30 | U | 31 | cWW | ncWWa | ncwW | cWW | |||
| 5 | IL_7ENI_001 | 7ENI | 0.1029 | 0 | B | sgRNA (74-MER) | A | 42 | A | 43 | U | 44 | C | 45 | U | 46 | * | A | 29 | C | 30 | U | 31 | cWW | ncSH | ncwW | cWW | |||
| 6 | IL_6DME_002 | 6DME | 0.3615 | 0 | 8-nt loop receptor | A | ppGpp Riboswitch | G | 15 | G | 16 | A | 17 | U | 18 | C | 19 | * | G | 37 | A | 38 | C | 39 | cWW | cWW | ncSH | ncSH | tHS | cWW |
| 7 | IL_6CK5_002 | 6CK5 | 0.4077 | 0 | 8-nt loop receptor | A | Guanidine-I riboswitch | G | 16 | G | 17 | U | 18 | U | 19 | C | 20 | * | G | 42 | A | 43 | C | 44 | cWW | cWW | cSH | cSH | tHS | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AAUCU*ACU | 2 |
| CGAAG*CAG | 1 |
| GACCC*GAC | 1 |
| AAGCU*ACU | 1 |
| GGAUC*GAC | 1 |
| GGUUC*GAC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AUC*C | 2 |
| GAA*A | 1 |
| ACC*A | 1 |
| AGC*C | 1 |
| GAU*A | 1 |
| GUU*A | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 2 parents |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (5)
- 8-nt loop receptor (2)
- Basepair signature
- cWW-cWW-L-cWW-L
- Heat map statistics
- Min 0.10 | Avg 0.42 | Max 0.98
Coloring options: