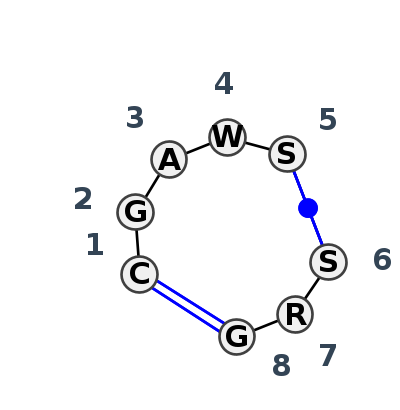
Motif IL_93336.2 Version IL_93336.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | break | 6 | 7 | 8 | 1-8 | 2-7 | 5-6 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_8CRE_410 | 8CRE | 0.0567 | 1 | CM | SSU rRNA | C | 374 | G | 375 | A | 376 | U | 377 | C | 379 | * | G | 361 | G | 362 | G | 363 | cWW | cWW | |
| 2 | IL_8GLP_212 | 8GLP | 0.0000 | 1 | S2 | SSU rRNA | C | 424 | G | 425 | A | 426 | U | 427 | C | 429 | * | G | 411 | G | 412 | G | 413 | cWW | cWW | |
| 3 | IL_8P9A_393 | 8P9A | 0.0638 | 1 | sR | SSU rRNA | C | 376 | G | 377 | A | 378 | U | 379 | C | 381 | * | G | 363 | G | 364 | G | 365 | cWW | cWW | |
| 4 | IL_9H3G_175 | 9H3G | 0.1372 | 1 | h1 | SSU rRNA | C | 378 | G | 379 | A | 380 | U | 381 | C | 383 | * | G | 365 | A | 366 | G | 367 | cWW | cWW | |
| 5 | IL_9E6Q_019 | 9E6Q | 0.5654 | 1 | 1 | LSU rRNA | C | 570 | G | 571 | A | 572 | A | 573 | G | 575 | * | C | 630 | A | 631 | G | 632 | cWW | tSH | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CGAUUC*GGG | 2 |
| CGAU(OMU)C*GGG | 1 |
| CGAUUC*GAG | 1 |
| CGAAAG*CAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAUU*G | 2 |
| GAU(OMU)*G | 1 |
| GAUU*A | 1 |
| GAAA*A | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (5)
- Basepair signature
- cWW-L-R-L-cWW-L
- Heat map statistics
- Min 0.06 | Avg 0.23 | Max 0.61
Coloring options: