- Structure title
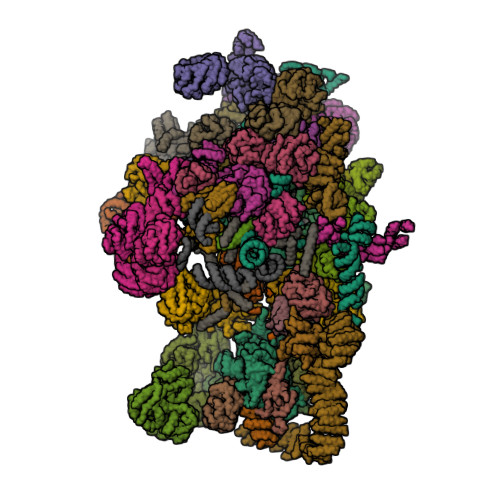
- Architecture of the yeast small subunit processome
- Authors
- Release date
- 2016-12-21
- Experimental technique
- ELECTRON MICROSCOPY
- Resolution
- 5.1 Å
- Chains
-
3 RNA chains from
Saccharomyces cerevisiae
-
3 DNA chains
-
-
52 other chains
- Nucleic acid compounds
5' external transcribed spacer, 18S ribosomal RNA, U3 snoRNA, 5' domain-associated, 3' domain-associated, Unassigned RNA helices
- Other compounds
rpS6_ES6, rpS4_ES4, rpS5_US7, rpS7_eS7, rpS8_eS8, rpS9_uS4, rpS16_uS9, rpS11_uS17, rpS22_uS8, rpS24_eS24, rpS28_eS28, Utp4, UtpA_CTD1, UtpA_CTD2, UtpA_CTD2, Beta-propeller 2, Utp17, Utp1, Utp6, Utp12, Utp13, Utp18, Utp21, Beta-propeller 5, Enp2, UtpA_CTD4, Kre33, Kre33, Imp3, Nop56, Nop58, Nop1, Nop1, Snu13, Snu13, Ribosomal RNA-processing protein 9, RNA 3'-terminal phosphate cyclase-like protein, Bms1,Ribosome biogenesis protein BMS1,Bms1, Ribosomal RNA small subunit methyltransferase NEP1, Ribosomal RNA small subunit methyltransferase NEP1, Utp24, Imp4, Utp30, Unassigned KH domain, Utp20, Repeat protein 2, 40S ribosomal protein S23-A, Beta-propeller 1, Beta-propeller 1, Beta-propeller 1, Repeat protein 1, Unassigned protein helices
Pairwise Interactions
Interactions annotated by
FR3D:
View all
RNA 3D Motifs
In the current release of the
RNA 3D Motif Atlas, 5TZS contains:
-
13 hairpin loops from 0 motif groups
-
34 internal loops from 0 motif groups
-
4 junction loops from 0 motif groups
More
3D Redundancy
5TZS has chains which belong to the following equivalence classes in the current representative set:
Similar structures
None found