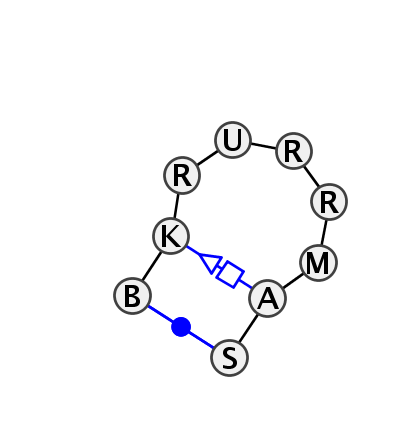
Motif HL_35608.1 Version HL_35608.1 of this group appears in releases 2.0 to 2.0
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 1-9 | 2-8 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_5J7L_192 | 5J7L | 0.1729 | 1 | DA | LSU rRNA | U | 2404 | G | 2405 | A | 2407 | U | 2408 | G | 2409 | G | 2410 | A | 2411 | A | 2412 | G | 2413 | cWW | tSH |
| 2 | HL_5FDU_058 | 5FDU | 0.1178 | 1 | 1A | LSU rRNA | C | 2416 | G | 2417 | G | 2419 | U | 2420 | G | 2421 | G | 2422 | A | 2423 | A | 2424 | G | 2425 | cWW | tSH |
| 3 | HL_4IOA_058 | 4IOA | 0.0000 | 1 | X | LSU rRNA | C | 2383 | G | 2384 | G | 2386 | U | 2387 | G | 2388 | G | 2389 | A | 2390 | A | 2391 | G | 2392 | cWW | ntSH |
| 4 | HL_3J9W_095 | 3J9W | 0.1422 | 1 | BA | LSU rRNA | C | 2433 | G | 2434 | A | 2436 | U | 2437 | G | 2438 | G | 2439 | A | 2440 | A | 2441 | G | 2442 | cWW | tSH |
| 5 | HL_5G2X_002 | 5G2X | 0.6997 | 3 | A | Group II catalytic intron | G | 48 | U | 49 | A | 53 | U | 54 | A | 55 | A | 56 | C | 57 | A | 58 | C | 59 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CGUGUGGAAG | 2 |
| UGAAUGGAAG | 1 |
| CGCAUGGAAG | 1 |
| GUAAGAUAACAC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GUGUGGAA | 2 |
| GAAUGGAA | 1 |
| GCAUGGAA | 1 |
| UAAGAUAACA | 1 |
Release history
| Release | 2.0 |
|---|---|
| Date | 2017-04-24 |
| Status | New id, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (5)
- Basepair signature
- cWW-tSH-R-R-R-R-R
- Heat map statistics
- Min 0.12 | Avg 0.32 | Max 0.83
Coloring options: