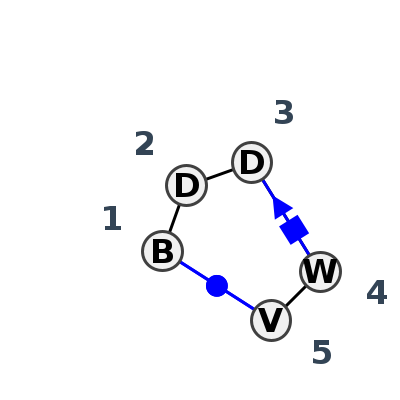
Motif HL_38901.2 Version HL_38901.2 of this group appears in releases 3.95 to 3.99
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 1-5 | 3-4 | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_5F9R_004 | 5F9R | 0.3894 | 0 | Other HL | A | tracrRNA | G | 105 | A | 106 | G | 107 | U | 108 | C | 109 | cWW | ncSH |
| 2 | HL_3RG5_005 | 3RG5 | 0.2355 | 1 | tRNA D-loop | B | Selenocysteine transfer RNA | G | 15 | U | 16 | G | 19 | U | 20 | C | 20|||A | cWW | cSH |
| 3 | HL_3W3S_001 | 3W3S | 0.1850 | 1 | tRNA D-loop | B | Selenocysteine transfer RNA | C | 15 | U | 16 | G | 19 | U | 20 | G | 20|||A | cWW | cSH |
| 4 | HL_3AM1_001 | 3AM1 | 0.0000 | 1 | tRNA D-loop | B | ASL-truncated tRNA | U | 18 | U | 19 | G | 21 | U | 22 | A | 23 | cWW | cSH |
| 5 | HL_3ADD_001 | 3ADD | 0.2334 | 1 | tRNA D-loop | C | Selenocysteine transfer RNA | G | 15 | U | 16 | G | 19 | U | 20 | C | 20|||A | cWW | cSH |
| 6 | HL_8P9A_219 | 8P9A | 0.6098 | 1 | sR | SSU rRNA | C | 1359 | A | 1360 | U | 1362 | U | 1363 | G | 1364 | cWW | ||
| 7 | HL_4KR9_004 | 4KR9 | 0.9092 | 1 | X | RNA (39-MER) | G | 19 | G | 20 | A | 22 | A | 23 | C | 24 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GUGGUC | 2 |
| GAGUC | 1 |
| CUGGUG | 1 |
| UUGGUA | 1 |
| CAUUUG | 1 |
| GGAAAC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UGGU | 4 |
| AGU | 1 |
| AUUU | 1 |
| GAAA | 1 |
- Annotations
-
- tRNA D-loop (4)
- (2)
- Other HL (1)
- Basepair signature
- cWW-F-cSH
- Heat map statistics
- Min 0.17 | Avg 0.47 | Max 0.92
Coloring options: