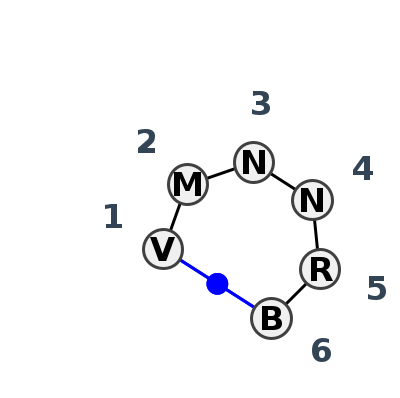
Motif HL_39454.2 Version HL_39454.2 of this group appears in releases 3.94 to 3.94
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 1-6 | 2-5 | 3-4 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_6WZR_001 | 6WZR | 0.4095 | 2 | A | ZMP/ZTP riboswitch | G | 28 | A | 29 | A | 30 | A | 33 | A | 34 | C | 35 | cWW | ||
| 2 | HL_5BTP_003 | 5BTP | 0.3997 | 2 | B | ZMP/ZTP riboswitch | G | 28 | A | 29 | A | 30 | A | 33 | A | 34 | C | 35 | cWW | ncsS | |
| 3 | HL_8SYK_002 | 8SYK | 0.1512 | 2 | A | RNA device 43 truncation mutant 3 (U100C) | C | 32 | C | 33 | A | 34 | U | 37 | A | 38 | G | 39 | cWW | ||
| 4 | HL_2OEU_002 | 2OEU | 0.0000 | 2 | A | Hammerhead Ribozyme | C | 25 | C | 26 | A | 27 | U | 30 | A | 31 | G | 32 | cWW | ||
| 5 | HL_9DFE_024 | 9DFE | 0.7891 | 2 | 1A | LSU rRNA | C | 884 | C | 885 | C | 886 | C | 889 | A | 890 | G | 892 | cWW | ncSW | ntSH |
| 6 | HL_8P9A_133 | 8P9A | 0.8151 | 3 | AR | LSU rRNA | G | 910 | C | 911 | G | 912 | A | 915 | A | 917 | C | 918 | cWW | cWW | tSW |
| 7 | HL_6CF2_001 | 6CF2 | 0.8571 | 2 | G | RNA (35-MER) | A | 13 | C | 14 | U | 15 | G | 18 | G | 19 | U | 20 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GAAUUAAC | 2 |
| CCAAAUAG | 2 |
| CCCACCAG | 1 |
| GCGAAAGAC | 1 |
| ACUUCGGU | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AAUUAA | 2 |
| CAAAUA | 2 |
| CCACCA | 1 |
| CGAAAGA | 1 |
| CUUCGG | 1 |
Release history
| Release | 3.94 |
|---|---|
| Date | 2025-02-26 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (7)
- Basepair signature
- cWW-F-F-F-F
- Heat map statistics
- Min 0.08 | Avg 0.55 | Max 0.92
Coloring options: