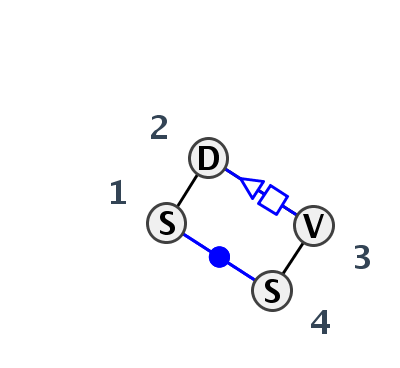
Motif HL_53845.2 Version HL_53845.2 of this group appears in releases 3.85 to 3.85
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 1-4 | 2-3 | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_7A0S_046 | 7A0S | 0.5812 | 3 | GNRA | X | LSU rRNA | C | 1797 | G | 1798 | A | 1802 | G | 1803 | cWW | tSH |
| 2 | HL_6N5P_002 | 6N5P | 0.5410 | 3 | A | cyclic di-AMP riboswitch | G | 58 | U | 59 | G | 63 | C | 64 | ncWW | ||
| 3 | HL_6CZR_074 | 6CZR | 0.5238 | 2 | GNRA | 1a | SSU rRNA | G | 153 | G | 154 | A | 157 | C | 158 | ncWW | ntSH |
| 4 | HL_1M5K_003 | 1M5K | 0.4521 | 2 | GNRA | B | Hairpin ribozyme | G | 74 | G | 75 | A | 78 | C | 79 | cWW | ntSH |
| 5 | HL_2CZJ_007 | 2CZJ | 0.4470 | 2 | UNCG variation | H | tmRNA | G | 25 | U | 26 | G | 29 | C | 30 | cWW | |
| 6 | HL_8C3A_206 | 8C3A | 0.0752 | 2 | GNRA variation | CM | SSU rRNA | C | 567 | A | 568 | C | 571 | G | 572 | cWW | |
| 7 | HL_4V88_200 | 4V88 | 0.0000 | 2 | GNRA variation | A6 | SSU rRNA | C | 569 | A | 570 | C | 573 | G | 574 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GGAAAC | 2 |
| CAGCCG | 2 |
| CGAACAG | 1 |
| GUGAAGC | 1 |
| GUUCGC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAAA | 2 |
| AGCC | 2 |
| GAACA | 1 |
| UGAAG | 1 |
| UUCG | 1 |
Release history
| Release | 3.85 |
|---|---|
| Date | 2024-06-19 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- GNRA (3)
- GNRA variation (2)
- UNCG variation (1)
- (1)
- Basepair signature
- cWW-tSH
- Heat map statistics
- Min 0.08 | Avg 0.39 | Max 0.71
Coloring options: