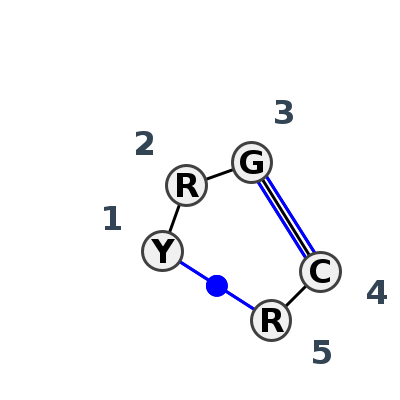
Motif IL_16386.3 Version IL_16386.3 of this group appears in releases 3.90 to 3.92
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | break | 4 | 5 | 1-5 | 3-4 | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_4R4V_005 | 4R4V | 0.9157 | 1 | Intercalated cWS | A | VS ribozyme RNA | U | 719 | A | 720 | G | 722 | * | C | 755 | A | 756 | cWW | cWW |
| 2 | IL_8HBA_003 | 8HBA | 0.0768 | 0 | B | RNA (55-MER) | C | 32 | G | 33 | G | 34 | * | C | 5 | G | 6 | cWW | cWW | |
| 3 | IL_8I3Z_001 | 8I3Z | 0.0795 | 0 | B+A | RNA (31-MER) | C | 32 | G | 33 | G | 34 | * | C | 5 | G | 6 | cWW | cWW | |
| 4 | IL_8GXB_001 | 8GXB | 0.0000 | 0 | A | NAD+ II riboswitch | C | 40 | G | 41 | G | 42 | * | C | 5 | G | 6 | cWW | cWW | |
| 5 | IL_8GXB_003 | 8GXB | 0.1032 | 0 | B | NAD+ II riboswitch | C | 40 | G | 41 | G | 42 | * | C | 5 | G | 6 | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CGG*CG | 4 |
| UAAG*CA | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| G* | 4 |
| AA* | 1 |
- Annotations
-
- (4)
- Intercalated cWS (1)
- Basepair signature
- cWW-L-cWW
- Heat map statistics
- Min 0.07 | Avg 0.34 | Max 0.95
Coloring options: