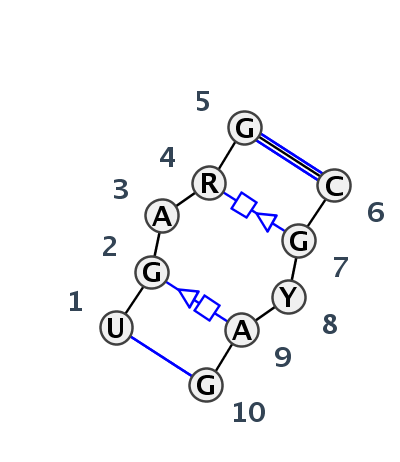
Motif IL_24484.1 Version IL_24484.1 of this group appears in releases 3.87 to 3.87
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | break | 6 | 7 | 8 | 9 | 10 | 1-10 | 2-9 | 3-8 | 4-7 | 5-6 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_7A0S_049 | 7A0S | 0.0000 | 0 | tSH-tHW-tHS | X | LSU rRNA | U | 1370 | G | 1371 | A | 1372 | G | 1373 | G | 1374 | * | C | 1383 | G | 1384 | C | 1385 | A | 1386 | G | 1387 | cWW | tSH | tHW | tHS | cWW |
| 2 | IL_8VTW_052 | 8VTW | 0.2489 | 0 | tSH-tHW-tHS (H) | 1A | LSU rRNA | U | 1357 | G | 1358 | A | 1359 | A | 1360 | G | 1361 | * | C | 1370 | G | 1371 | U | 1372 | A | 1373 | G | 1374 | cWW | tSH | tWW | tHS | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UGAGG*CGCAG | 1 |
| UGAAG*CGUAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAG*GCA | 1 |
| GAA*GUA | 1 |
Release history
| Release | 3.87 |
|---|---|
| Date | 2024-08-14 |
| Status | New id, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- tSH-tHW-tHS (1)
- tSH-tHW-tHS (H) (1)
- Basepair signature
- cWW-tSH-L-R-tHS-cWW
- Heat map statistics
- Min 0.25 | Avg 0.12 | Max 0.25
Coloring options: