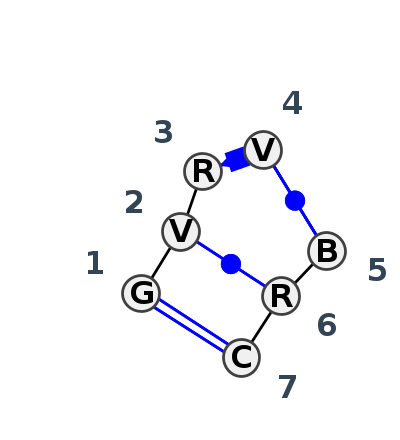
Motif IL_63450.5 Version IL_63450.5 of this group appears in releases 3.89 to 3.90
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | break | 5 | 6 | 7 | 1-7 | 2-6 | 3-4 | 3-6 | 4-5 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_4V9F_072 | 4V9F | 0.3975 | 0 | 0 | LSU rRNA | G | 2090 | G | 2091 | G | 2092 | G | 2093 | * | U | 2652 | A | 2653 | C | 2654 | cWW | cWW | cSH | cWW | ||
| 2 | IL_6DME_003 | 6DME | 0.3723 | 0 | A | ppGpp Riboswitch | G | 33 | A | 34 | A | 35 | G | 36 | * | C | 20 | A | 21 | C | 22 | cWW | cwW | cSH | cWW | ||
| 3 | IL_7SZU_003 | 7SZU | 0.3118 | 0 | R | RNA aptamer | G | 49 | C | 50 | A | 51 | C | 52 | * | G | 25 | A | 26 | C | 27 | cWW | cWW | cSH | cWW | ||
| 4 | IL_5TBW_091 | 5TBW | 0.0941 | 0 | Major groove platform with extra pair | 1 | LSU rRNA | G | 2391 | C | 2392 | G | 2393 | G | 2394 | * | U | 2986 | A | 2987 | C | 2988 | cWW | cWW | cSH | cWW | |
| 5 | IL_8C3A_096 | 8C3A | 0.0000 | 0 | Major groove platform with extra pair (H) | 1 | LSU rRNA | G | 2369 | C | 2370 | G | 2371 | G | 2372 | * | U | 2958 | A | 2959 | C | 2960 | cWW | cWW | cSH | cWW | |
| 6 | IL_5TBW_087 | 5TBW | 0.3962 | 1 | Major groove platform with extra pair | 1 | LSU rRNA | G | 2206 | A | 2207 | A | 2208 | G | 2210 | * | C | 2235 | G | 2236 | C | 2237 | cWW | cWW | |||
| 7 | IL_3DD2_004 | 3DD2 | 0.5989 | 2 | B | RNA (26-MER) | G | 3 | A | 4 | A | 5 | A | 8 | * | UFT | 21 | A | 22 | CFL | 23 | cWW | cSW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GCGG*UAC | 2 |
| GGGG*UAC | 1 |
| GAAG*CAC | 1 |
| GCAC*GAC | 1 |
| GAAUG*CGC | 1 |
| GAA(CFL)AA*(UFT)A(CFL) | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| CG*A | 2 |
| GG*A | 1 |
| AA*A | 1 |
| CA*A | 1 |
| AAU*G | 1 |
| AA(CFL)A*A | 1 |
- Annotations
-
- (4)
- Major groove platform with extra pair (2)
- Major groove platform with extra pair (H) (1)
- Basepair signature
- cWW-cHS-cWW-cWW-cWW
- Heat map statistics
- Min 0.09 | Avg 0.37 | Max 0.64
Coloring options: