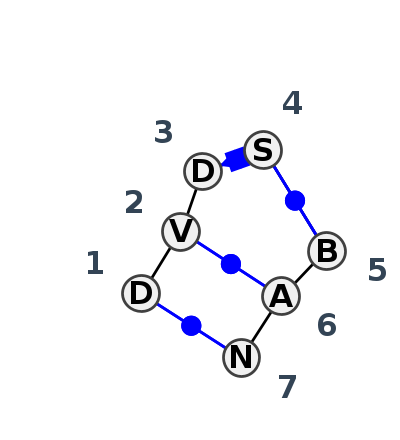
Motif IL_75527.3 Version IL_75527.3 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | break | 5 | 6 | 7 | 1-7 | 2-4 | 2-6 | 3-4 | 3-6 | 4-5 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_6CK5_011 | 6CK5 | 0.7782 | 2 | A | Guanidine-I riboswitch | U | 36 | G | 37 | U | 39 | G | 41 | * | C | 21 | A | 22 | G | 23 | cWW | cWH | cWW | cWW | |||
| 2 | IL_4RUM_002 | 4RUM | 0.4918 | 1 | A | NiCo riboswitch | U | 38 | G | 39 | A | 41 | G | 42 | * | C | 20 | A | 21 | A | 22 | cWW | cWH | cWW | ||||
| 3 | IL_9AXU_085 | 9AXU | 0.0799 | 0 | 2 | LSU rRNA | G | 2479 | C | 2480 | G | 2481 | G | 2482 | * | U | 3081 | A | 3082 | C | 3083 | cWW | cWW | cSH | cWW | |||
| 4 | IL_9H3G_100 | 9H3G | 0.0958 | 0 | A | LSU rRNA | OMG | 2389 | C | 2390 | G | 2391 | G | 2392 | * | U | 2985 | A | 2986 | C | 2987 | cWW | cWW | cSH | cWW | |||
| 5 | IL_8GLP_127 | 8GLP | 0.0793 | 0 | L5 | LSU rRNA | G | 3895 | C | 3896 | G | 3897 | G | 3898 | * | U | 4563 | A | 4564 | C | 4565 | cWW | cWW | cSH | cWW | |||
| 6 | IL_8P9A_319 | 8P9A | 0.0560 | 0 | Major groove platform with extra pair (H) | AR | LSU rRNA | G | 2391 | C | 2392 | G | 2393 | G | 2394 | * | U | 2986 | A | 2987 | C | 2988 | cWW | cWW | cSH | cWW | ||
| 7 | IL_8OI5_093 | 8OI5 | 0.0000 | 0 | 1 | LSU rRNA | G | 2369 | C | 2370 | G | 2371 | G | 2372 | * | U | 2958 | A | 2959 | C | 2960 | cWW | cWW | cSH | cWW | |||
| 8 | IL_7SZU_003 | 7SZU | 0.3142 | 0 | R | RNA aptamer | G | 49 | C | 50 | A | 51 | C | 52 | * | G | 25 | A | 26 | C | 27 | cWW | cWW | cSH | cWW | |||
| 9 | IL_9AXU_061 | 9AXU | 0.5637 | 0 | 2 | LSU rRNA | A | 1653 | A | 1654 | G | 1655 | G | 1656 | * | C | 1879 | A | 1880 | U | 1881 | cWW | ncWw | cSH | cWW | |||
| 10 | IL_8B0X_178 | 8B0X | 0.4160 | 0 | A | SSU rRNA | G | 1491 | A | 1492 | A | 1493 | G | 1494 | * | 5MC | 1407 | A | 1408 | C | 1409 | cWW | cwW | cSH | cWW | |||
| 11 | IL_6DME_003 | 6DME | 0.3817 | 0 | A | ppGpp Riboswitch | G | 33 | A | 34 | A | 35 | G | 36 | * | C | 20 | A | 21 | C | 22 | cWW | cwW | cSH | cWW | |||
| 12 | IL_4V9F_072 | 4V9F | 0.4069 | 0 | 0 | LSU rRNA | G | 2090 | G | 2091 | G | 2092 | G | 2093 | * | U | 2652 | A | 2653 | C | 2654 | cWW | cWW | cSH | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GCGG*UAC | 4 |
| UGUUCG*CAG | 1 |
| UGCAG*CAA | 1 |
| (OMG)CGG*UAC | 1 |
| GCAC*GAC | 1 |
| AAGG*CAU | 1 |
| GAAG*(5MC)AC | 1 |
| GAAG*CAC | 1 |
| GGGG*UAC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| CG*A | 5 |
| AA*A | 2 |
| GUUC*A | 1 |
| GCA*A | 1 |
| CA*A | 1 |
| AG*A | 1 |
| GG*A | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 2 parents |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (11)
- Major groove platform with extra pair (H) (1)
- Basepair signature
- cWW-cWW-cSH-cWW
- Heat map statistics
- Min 0.03 | Avg 0.41 | Max 0.91
Coloring options: