- Structure title
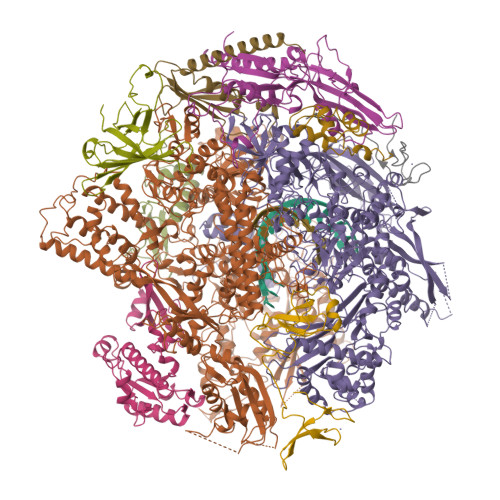
- RNA POLYMERASE II ELONGATION COMPLEX
- Authors
- Gnatt, A.L., Cramer, P., Kornberg, R.D.
- Release date
- 2001-04-23
- Experimental technique
- X-RAY DIFFRACTION
- Resolution
- 3.3 Å
- Chains
-
1 RNA chain from
unknown source
-
1 DNA chain
-
-
10 other chains
- Nucleic acid compounds
5'-R(P*GP*AP*CP*CP*AP*GP*GP*CP*A)-3', 5'-D(P*AP*AP*AP*TP*GP*CP*CP*TP*GP*GP*TP*CP*T)-3'
- Other compounds
DNA-DIRECTED RNA POLYMERASE II LARGEST SUBUNIT, DNA-DIRECTED RNA POLYMERASE II 140KD POLYPEPTIDE, DNA-DIRECTED RNA POLYMERASE II 45KD POLYPEPTIDE, DNA-DIRECTED RNA POLYMERASE II 27KD POLYPEPTIDE, DNA-DIRECTED RNA POLYMERASE II 23KD POLYPEPTIDE, DNA-DIRECTED RNA POLYMERASE II 14.5KD POLYPEPTIDE, DNA-DIRECTED RNA POLYMERASE II 14.2KD POLYPEPTIDE, DNA-DIRECTED RNA POLYMERASE II 8.3KD POLYPEPTIDE, DNA-DIRECTED RNA POLYMERASE II 13.6KD POLYPEPTIDE, DNA-DIRECTED RNA POLYMERASE II 7.7KD POLYPEPTIDE
Pairwise Interactions
Interactions annotated by
FR3D:
View all
RNA 3D Motifs
In the current release of the
RNA 3D Motif Atlas, 1I6H contains:
-
0 hairpin loops from 0 motif groups
-
1 internal loop from 0 motif groups
-
0 junction loops from 0 motif groups
More
3D Redundancy
This PDB is not included in any representative set equivalence class. Possibly, this PDB doesn't contain any complete nucleotides.
Similar structures
None found