- Structure title
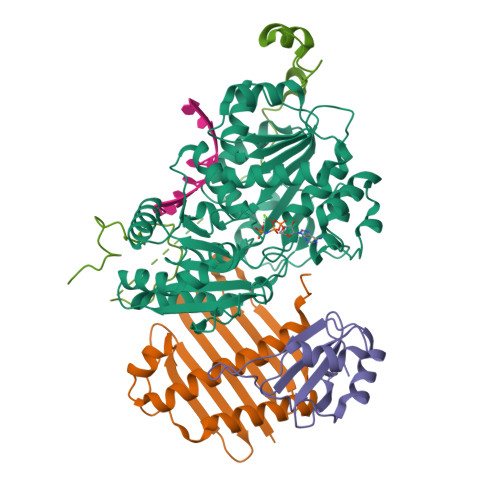
- The crystal structure of the Exon Junction Complex at 3.2 A resolution
- Authors
- Bono, F., Ebert, J., Lorentzen, E., Conti, E.
- Release date
- 2006-08-30
- Experimental technique
- X-RAY DIFFRACTION
- Resolution
- 3.2 Å
- Chains
-
2 RNA chains from
unknown source
-
8 other chains
- Nucleic acid compounds
5'-R(*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*U)-3', 5'-R(*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*UP*U)-3'
- Other compounds
ATP-DEPENDENT RNA HELICASE DDX48, ATP-DEPENDENT RNA HELICASE DDX48, PROTEIN MAGO NASHI HOMOLOG, RNA-BINDING PROTEIN 8A, PROTEIN MAGO NASHI HOMOLOG, RNA-BINDING PROTEIN 8A, PROTEIN CASC3, PROTEIN CASC3
Pairwise Interactions
Interactions annotated by
FR3D:
View all
RNA 3D Motifs
In the current release of the
RNA 3D Motif Atlas, 2J0Q contains:
-
0 hairpin loops from 0 motif groups
-
0 internal loops from 0 motif groups
-
0 junction loops from 0 motif groups
More
3D Redundancy
2J0Q has chains which belong to the following equivalence classes in the current representative set:
Similar structures
None found