- Structure title
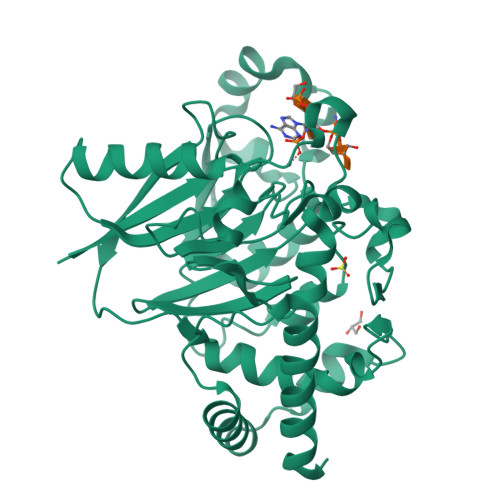
- Dbr1 in complex with synthetic linear RNA
- Authors
- Montemayor, E.J., Katolik, A., Clark, N.E., Taylor, A.B., Schuermann, J.P., Combs, D.J., Johnsson, R., Holloway, S.P., Stevens, S.W., Damha, M.J., Hart, P.J.
- Release date
- 2014-08-27
- Experimental technique
- X-RAY DIFFRACTION
- Resolution
- 2.1 Å
- Chains
-
5 RNA chains from
synthetic construct
-
5 other chains
- Nucleic acid compounds
RNA (5'-R(*CP*UP*AP*(A2P)P*AP*CP*AP*A)-3'), RNA (5'-R(*CP*UP*AP*(A2P)P*AP*CP*AP*A)-3'), RNA (5'-R(*CP*UP*AP*(A2P)P*AP*CP*AP*A)-3'), RNA (5'-R(*CP*UP*AP*(A2P)P*AP*CP*AP*A)-3'), RNA (5'-R(*CP*UP*AP*(A2P)P*AP*CP*AP*A)-3')
- Other compounds
RNA lariat debranching enzyme, putative, RNA lariat debranching enzyme, putative, RNA lariat debranching enzyme, putative, RNA lariat debranching enzyme, putative, RNA lariat debranching enzyme, putative
Pairwise Interactions
Interactions annotated by
FR3D:
View all
RNA 3D Motifs
In the current release of the
RNA 3D Motif Atlas, 4PEH contains:
-
0 hairpin loops from 0 motif groups
-
0 internal loops from 0 motif groups
-
0 junction loops from 0 motif groups
More
3D Redundancy
4PEH has chains which belong to the following equivalence classes in the current representative set:
Similar structures
None found