- Structure title
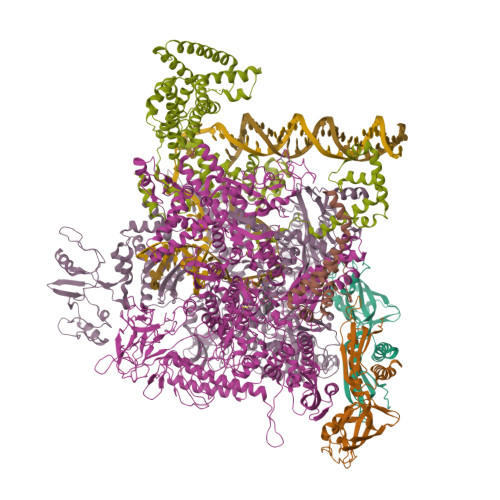
- E. coli Transcription Initiation Complex - 16-bp spacer and 4-nt RNA
- Authors
- Release date
- 2015-04-22
- Experimental technique
- X-RAY DIFFRACTION
- Resolution
- 6.0 Å
- Chains
-
3 RNA chains from
synthetic construct
-
6 DNA chains
-
-
18 other chains
- Nucleic acid compounds
RNA (5'-D(*(GTP))-R(P*AP*GP*U)-3'), RNA (5'-D(*(GTP))-R(P*AP*GP*U)-3'), RNA (5'-D(*(GTP))-R(P*AP*GP*U)-3'), NT strand DNA (49-MER), T strand DNA (49-MER), NT strand DNA (49-MER), T strand DNA (49-MER), NT strand DNA (49-MER), T strand DNA (49-MER)
- Other compounds
DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, DNA-directed RNA polymerase subunit beta', DNA-directed RNA polymerase subunit omega, RNA polymerase sigma factor RpoD, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, DNA-directed RNA polymerase subunit beta', DNA-directed RNA polymerase subunit omega, RNA polymerase sigma factor RpoD, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, DNA-directed RNA polymerase subunit beta', DNA-directed RNA polymerase subunit omega, RNA polymerase sigma factor RpoD
Pairwise Interactions
Interactions annotated by
FR3D:
View all
RNA 3D Motifs
In the current release of the
RNA 3D Motif Atlas, 4YLO contains:
-
0 hairpin loops from 0 motif groups
-
22 internal loops from 0 motif groups
-
0 junction loops from 0 motif groups
More
3D Redundancy
4YLO has chains which belong to the following equivalence classes in the current representative set:
Similar structures
None found