- Structure title
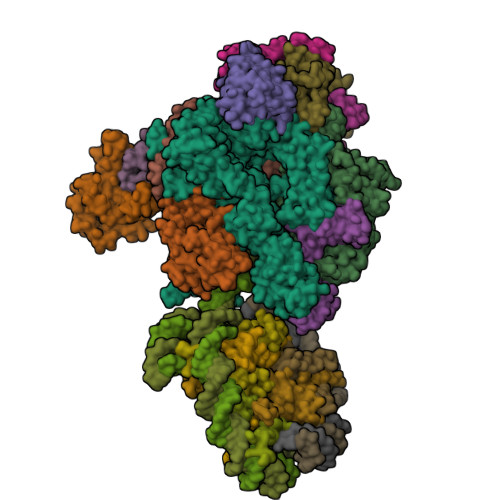
- RNA polymerase II elongation complex stalled at SHL(-1) of the nucleosome, with foreign DNA
- Authors
- Release date
- 2018-10-03
- Experimental technique
- ELECTRON MICROSCOPY
- Resolution
- 5.6 Å
- Chains
-
1 RNA chain from
synthetic construct
-
4 DNA chains
-
-
20 other chains
- Nucleic acid compounds
RNA (5'-R(P*GP*GP*UP*GP*UP*CP*UP*UP*GP*GP*G)-3'), DNA (42-MER), DNA (42-MER), DNA (198-MER), DNA (198-MER)
- Other compounds
DNA-directed RNA polymerase subunit, DNA-directed RNA polymerase subunit beta, RNA polymerase II third largest subunit B44, part of central core, RNA polymerase II subunit B32, RNA polymerase subunit ABC27, common to RNA polymerases I, II, and III, RNA polymerase subunit ABC23, common to RNA polymerases I, II, and III, RNA polymerase II subunit, DNA-directed RNA polymerases I, II, and III subunit RPABC3, DNA-directed RNA polymerase subunit, RNA polymerase subunit ABC10-beta, common to RNA polymerases I, II, and III, RNA polymerase II subunit B12.5, RNA polymerase subunit ABC10-alpha, Histone H3.3, Histone H4, Histone H2A type 1-B/E, Histone H2B type 1-J, Histone H3.3, Histone H4, Histone H2A type 1-B/E, Histone H2B type 1-J
Pairwise Interactions
Interactions annotated by
FR3D:
View all
RNA 3D Motifs
In the current release of the
RNA 3D Motif Atlas, 6A5L contains:
-
0 hairpin loops from 0 motif groups
-
32 internal loops from 0 motif groups
-
0 junction loops from 0 motif groups
More
3D Redundancy
This PDB is not included in any representative set equivalence class. Possibly, this PDB doesn't contain any complete nucleotides.
Similar structures
None found