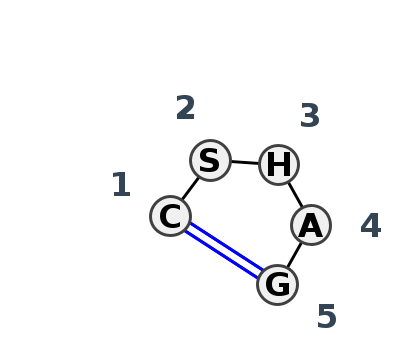
Motif HL_36488.1 Version HL_36488.1 of this group appears in releases 4.1 to 4.6
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 1-5 | 2-4 | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_4C9D_002 | 4C9D | 0.2318 | 0 | D | R3 REPEAT RNA CLEAVAGE PRODUCT | C | 21 | G | 22 | U | 23 | A | 24 | G | 25 | ncWW | |
| 2 | HL_4V9F_007 | 4V9F | 0.0000 | 2 | 0 | LSU rRNA | C | 195 | G | 196 | A | 198 | A | 199 | G | 201 | cWW | tSH |
| 3 | HL_8GLP_112 | 8GLP | 0.6281 | 1 | S2 | SSU rRNA | C | 1417 | C | 1418 | C | 1419 | A | 1421 | G | 1422 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CGUAG | 1 |
| CGCAACG | 1 |
| CCCGAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GUA | 1 |
| GCAAC | 1 |
| CCGA | 1 |
- Annotations
-
- (3)
- Basepair signature
- cWW-F-F-F
- Heat map statistics
- Min 0.23 | Avg 0.34 | Max 0.67
Coloring options: