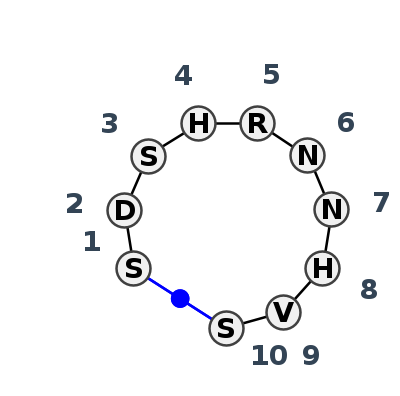
Motif HL_67407.2 Version HL_67407.2 of this group appears in releases 3.91 to 3.94
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 1-10 | 2-9 | 6-7 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_6WZR_002 | 6WZR | 0.4170 | 1 | A | ZMP/ZTP riboswitch | C | 62 | G | 63 | C | 65 | C | 66 | G | 67 | U | 68 | C | 69 | U | 70 | G | 71 | G | 72 | cWW | ||
| 2 | HL_5BTP_004 | 5BTP | 0.4030 | 1 | B | ZMP/ZTP riboswitch | C | 62 | G | 63 | C | 65 | C | 66 | G | 67 | U | 68 | C | 69 | U | 70 | G | 71 | G | 72 | cWW | ||
| 3 | HL_9BZC_003 | 9BZC | 0.0000 | 1 | A | RNA (87-MER) | C | 77 | G | 78 | C | 80 | C | 81 | G | 82 | U | 83 | C | 84 | U | 85 | G | 86 | G | 87 | cWW | ||
| 4 | HL_4XWF_002 | 4XWF | 0.1542 | 1 | A | ZMP/ZTP riboswitch | C | 50 | G | 51 | C | 53 | C | 54 | G | 55 | C | 56 | C | 57 | U | 58 | G | 59 | G | 60 | cWW | ||
| 5 | HL_4ZNP_002 | 4ZNP | 0.2023 | 1 | A | ZMP/ZTP riboswitch | C | 58 | G | 59 | C | 61 | C | 62 | G | 63 | C | 64 | C | 65 | U | 66 | G | 67 | G | 68 | cWW | ||
| 6 | HL_4PJO_001 | 4PJO | 0.3238 | 1 | 1 | U1 RNA variant (48-MER) with 4-helix junction replaced by kissing loop (HIV-1 (Mal) DIS) and shorter stem-loop 4. | G | 17 | A | 18 | G | 20 | U | 21 | G | 22 | C | 23 | A | 24 | C | 25 | A | 26 | C | 27 | cWW | ntWW | |
| 7 | HL_7JRS_002 | 7JRS | 0.4331 | 1 | B | RNA 3D nanocage | G | 63 | A | 64 | G | 66 | C | 67 | A | 68 | G | 69 | G | 70 | C | 71 | A | 72 | C | 73 | cWW | ncwW | |
| 8 | HL_7JRS_004 | 7JRS | 0.7701 | 1 | A | RNA 3D nanocage | G | 63 | A | 64 | G | 66 | C | 67 | A | 68 | G | 69 | G | 70 | C | 71 | A | 72 | C | 73 | ncWW | tWW | |
| 9 | HL_7A0S_002 | 7A0S | 0.7321 | 1 | X | LSU rRNA | C | 85 | U | 86 | G | 88 | A | 89 | G | 90 | A | 91 | U | 92 | A | 93 | C | 94 | G | 95 | cWW | ncWW | ncSH |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CGACCGUCUGG | 3 |
| GAAGCAGGCAC | 2 |
| CGCCCGCCUGG | 1 |
| CGACCGCCUGG | 1 |
| GAGGUGCACAC | 1 |
| CUGGAGAUACG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GACCGUCUG | 3 |
| AAGCAGGCA | 2 |
| GCCCGCCUG | 1 |
| GACCGCCUG | 1 |
| AGGUGCACA | 1 |
| UGGAGAUAC | 1 |
- Annotations
-
- (9)
- Basepair signature
- cWW-F-F-F-F-F-F-F-F
- Heat map statistics
- Min 0.13 | Avg 0.51 | Max 0.81
Coloring options: