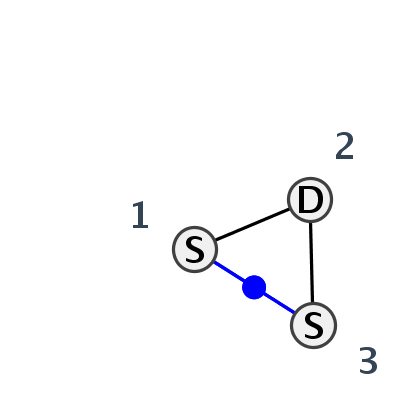
Motif HL_82739.1 Version HL_82739.1 of this group appears in releases 3.77 to 3.79
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 1-3 | |||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_2BH2_002 | 2BH2 | 0.3223 | 6 | D | 23S RIBOSOMAL RNA 1932-1968 | G | 1949 | G | 1950 | C | 1957 | cWW |
| 2 | HL_7A0S_011 | 7A0S | 0.3303 | 2 | X | LSU rRNA | C | 330 | U | 331 | G | 334 | cWW |
| 3 | HL_7A0S_003 | 7A0S | 0.3416 | 3 | X | LSU rRNA | G | 121 | G | 122 | C | 126 | cWW |
| 4 | HL_2ZNI_001 | 2ZNI | 0.3482 | 4 | C | bacterial tRNA | C | 13 | G | 14 | G | 21 | cWW |
| 5 | HL_4XWF_001 | 4XWF | 0.2456 | 3 | A | ZMP/ZTP riboswitch | G | 24 | G | 25 | C | 29 | cWW |
| 6 | HL_5J7L_143 | 5J7L | 0.2914 | 2 | DA | LSU rRNA | G | 319 | A | 320 | C | 323 | cWW |
| 7 | HL_7RQB_011 | 7RQB | 0.3127 | 2 | 1A | LSU rRNA | C | 319 | A | 320 | G | 323 | cWW |
| 8 | HL_4WF9_011 | 4WF9 | 0.3040 | 2 | X | LSU rRNA | C | 362 | A | 363 | G | 366 | cWW |
| 9 | HL_7RQB_003 | 7RQB | 0.2313 | 3 | 1A | LSU rRNA | G | 123 | G | 124 | C | 128 | cWW |
| 10 | HL_5J7L_136 | 5J7L | 0.1951 | 3 | DA | LSU rRNA | G | 123 | G | 124 | C | 128 | cWW |
| 11 | HL_4WF9_003 | 4WF9 | 0.1691 | 3 | X | LSU rRNA | G | 122 | G | 123 | C | 127 | cWW |
| 12 | HL_2QUS_003 | 2QUS | 0.0835 | 3 | A | Hammerhead ribozyme (type III) | C | 42 | G | 43 | G | 47 | cWW |
| 13 | HL_2QUW_003 | 2QUW | 0.0984 | 3 | B | Hammerhead ribozyme, cleaved fragment | C | 42 | G | 43 | G | 47 | cWW |
| 14 | HL_1WZ2_007 | 1WZ2 | 0.0000 | 3 | D | tRNA | C | 950 | G | 951 | G | 955 | cWW |
| 15 | HL_4KR9_002 | 4KR9 | 0.2430 | 3 | M | RNA (39-MER) | G | 19 | G | 20 | C | 24 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GGAAAC | 5 |
| CGUGAG | 2 |
| GGUAAGUUC | 1 |
| CUCAG | 1 |
| CGAAUAG | 1 |
| GAUAC | 1 |
| CAGAG | 1 |
| CAAAG | 1 |
| GGGAAC | 1 |
| CGUAGG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAAA | 5 |
| GUGA | 2 |
| GUAAGUU | 1 |
| UCA | 1 |
| GAAUA | 1 |
| AUA | 1 |
| AGA | 1 |
| AAA | 1 |
| GGAA | 1 |
| GUAG | 1 |
Coloring options: