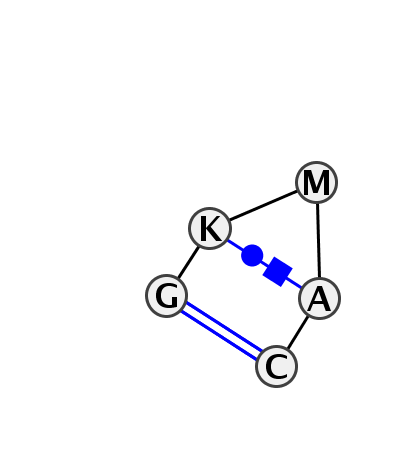
Motif HL_86069.1 Version HL_86069.1 of this group appears in releases 3.54 to 3.76
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 1-5 | 2-4 | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_3D2V_001 | 3D2V | 0.0000 | 1 | A | TPP riboswitch (THI element) | G | 17 | U | 19 | C | 20 | A | 21 | C | 22 | cWW | cWH |
| 2 | HL_7RQB_004 | 7RQB | 0.8812 | 1 | 1A | LSU rRNA | G | 139 | G | 140 | A | 141 | A | 142 | C | 142|||A | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GUUCAC | 1 |
| GGGAAC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UUCA | 1 |
| GGAA | 1 |
Release history
| Release | 3.54 | 3.55 | 3.56 | 3.57 | 3.58 | 3.59 | 3.60 | 3.61 | 3.62 | 3.63 | 3.64 | 3.65 | 3.66 | 3.67 | 3.68 | 3.69 | 3.70 | 3.71 | 3.72 | 3.73 | 3.74 | 3.75 | 3.76 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Date | 2022-02-02 | 2022-03-02 | 2022-03-30 | 2022-04-27 | 2022-05-25 | 2022-06-22 | 2022-07-20 | 2022-08-17 | 2022-10-12 | 2022-11-09 | 2022-12-07 | 2023-01-04 | 2023-02-01 | 2023-03-01 | 2023-03-29 | 2023-04-26 | 2023-05-24 | 2023-06-21 | 2023-07-19 | 2023-08-16 | 2023-09-13 | 2023-10-11 | 2023-11-08 |
| Status | New id, 1 parent | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (2)
- Basepair signature
- cWW-cWH-R
- Heat map statistics
- Min 0.88 | Avg 0.44 | Max 0.88
Coloring options: