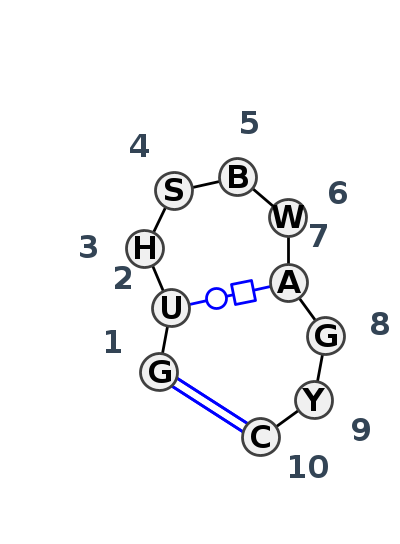
Motif HL_98864.4 Version HL_98864.4 of this group appears in releases 4.4 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 1-10 | 2-7 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_4RGE_003 | 4RGE | 0.1043 | 0 | T-loop with unstacked turn | C | type-P1 twister ribozyme | G | 29 | U | 30 | C | 31 | C | 32 | U | 33 | A | 34 | A | 35 | G | 36 | C | 37 | C | 38 | cWW | tWH |
| 2 | HL_4OJI_002 | 4OJI | 0.0000 | 0 | T-loop with unstacked turn | A | type-P1 twister ribozyme | G | 23 | U | 24 | C | 25 | C | 26 | C | 27 | A | 28 | A | 29 | G | 30 | C | 31 | C | 32 | cWW | tWH |
| 3 | HL_4QJD_004 | 4QJD | 0.2048 | 0 | T-loop with unstacked turn | D | type-P1 twister ribozyme | G | 43 | U | 44 | C | 45 | C | 46 | C | 47 | A | 48 | A | 49 | G | 50 | C | 51 | C | 52 | cWW | ntWW |
| 4 | HL_8UIW_002 | 8UIW | 0.6776 | 0 | T-loop with unstacked turn | N | azaaromatic riboswitch aptamer (yjdF RNA) | G | 50 | U | 51 | A | 52 | G | 53 | G | 54 | U | 55 | A | 56 | G | 57 | U | 58 | C | 59 | cWW | tWH |
| 5 | HL_9EBV_001 | 9EBV | 0.7932 | 0 | T-loop with unstacked turn (H) | A | azaaromatic riboswitch aptamer (yjdF RNA) | G | 21 | U | 22 | U | 23 | G | 24 | G | 25 | U | 26 | A | 27 | G | 28 | U | 29 | C | 30 | cWW | tWH |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GUCCCAAGCC | 2 |
| GUCCUAAGCC | 1 |
| GUAGGUAGUC | 1 |
| GUUGGUAGUC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UCCCAAGC | 2 |
| UCCUAAGC | 1 |
| UAGGUAGU | 1 |
| UUGGUAGU | 1 |
- Annotations
-
- T-loop with unstacked turn (4)
- T-loop with unstacked turn (H) (1)
- Basepair signature
- cWW-tWH-F-F-F-F-F-F
- Heat map statistics
- Min 0.10 | Avg 0.46 | Max 0.81
Coloring options: