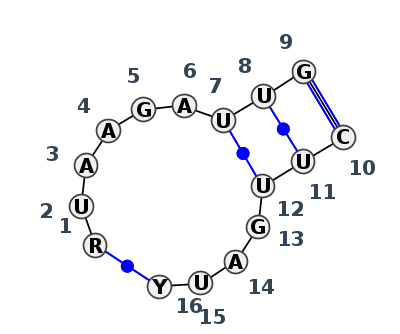
Motif IL_06668.1 Version IL_06668.1 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | break | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 1-16 | 3-14 | 5-14 | 6-13 | 7-12 | 8-11 | 9-10 | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_8P9A_470 | 8P9A | 0.0000 | 2 | Kink-turn (H) | sR | SSU rRNA | A | 1224 | U | 1225 | A | 1226 | A | 1227 | G | 1229 | A | 1230 | U | 1231 | U | 1232 | G | 1233 | * | C | 1252 | U | 1253 | U | 1254 | G | 1255 | A | 1256 | U | 1258 | U | 1259 | cWW | ntSs | cWw | cWw | cWW | ||
| 2 | IL_9H3G_216 | 9H3G | 0.3053 | 2 | h1 | SSU rRNA | G | 1225 | U | 1226 | A | 1227 | A | 1228 | G | 1230 | A | 1231 | OMU | 1232 | U | 1233 | G | 1234 | * | C | 1253 | U | 1254 | U | 1255 | G | 1256 | A | 1257 | U | 1259 | C | 1260 | cWW | ntsS | tSH | tHS | ncWw | ncWw | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AUAAGGAUUG*CUUGAUUU | 1 |
| GUAAGGA(OMU)UG*CUUGAUUC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UAAGGAUU*UUGAUU | 1 |
| UAAGGA(OMU)U*UUGAUU | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | New id, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- Kink-turn (H) (1)
- (1)
- Basepair signature
- cWW-L-R-L-R-L-R-L-cWW-L-cWW-cWW
- Heat map statistics
- Min 0.31 | Avg 0.15 | Max 0.31
Coloring options: