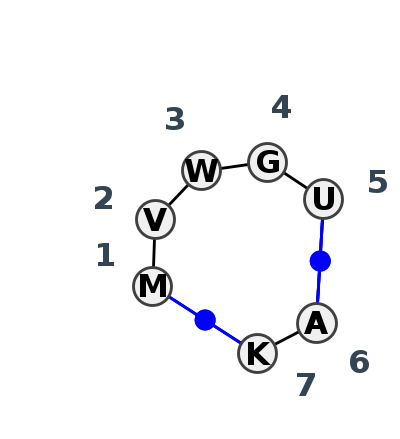
Motif IL_10396.2 Version IL_10396.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | break | 6 | 7 | 1-7 | 4-5 | 5-6 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_9E6Q_010 | 9E6Q | 0.2369 | 1 | 1 | LSU rRNA | A | 281 | G | 282 | A | 283 | G | 285 | U | 286 | * | A | 258 | U | 259 | cWW | cWW | |
| 2 | IL_4V9F_009 | 4V9F | 0.2187 | 1 | 0 | LSU rRNA | C | 260 | A | 261 | A | 262 | G | 264 | U | 265 | * | A | 248 | G | 249 | cWW | cWW | |
| 3 | IL_8GLP_008 | 8GLP | 0.1368 | 1 | L5 | LSU rRNA | A | 147 | C | 148 | A | 149 | G | 151 | U | 152 | * | A | 121 | U | 122 | cWW | cWW | |
| 4 | IL_9AXU_006 | 9AXU | 0.0000 | 1 | 2 | LSU rRNA | A | 150 | G | 151 | U | 152 | G | 154 | U | 155 | * | A | 123 | U | 124 | cWW | cWW | |
| 5 | IL_8P9A_236 | 8P9A | 0.0786 | 1 | AR | LSU rRNA | A | 144 | G | 145 | U | 146 | G | 148 | U | 149 | * | A | 123 | U | 124 | cWW | cSH | cWW |
| 6 | IL_8OI5_007 | 8OI5 | 0.0686 | 1 | 1 | LSU rRNA | A | 143 | G | 144 | U | 145 | G | 147 | U | 148 | * | A | 122 | U | 123 | cWW | cWW | |
| 7 | IL_9H3G_011 | 9H3G | 0.0893 | 1 | A | LSU rRNA | A | 142 | A | 143 | OMU | 144 | G | 146 | U | 147 | * | A | 121 | U | 122 | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AGUUGU*AU | 3 |
| AGAUGU*AU | 1 |
| CAAUGU*AG | 1 |
| ACAUGU*AU | 1 |
| AA(OMU)UGU*AU | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GUUG* | 3 |
| GAUG* | 1 |
| AAUG* | 1 |
| CAUG* | 1 |
| A(OMU)UG* | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (7)
- Basepair signature
- cWW-L-cWW-L-L
- Heat map statistics
- Min 0.06 | Avg 0.14 | Max 0.28
Coloring options: