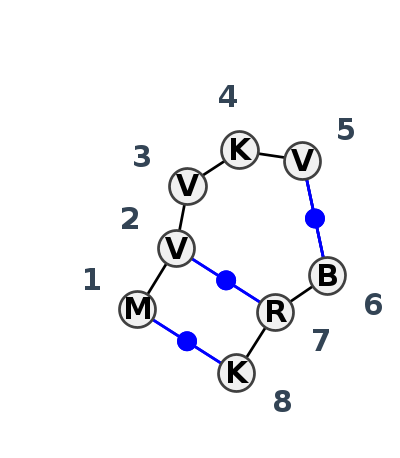
Motif IL_19668.2 Version IL_19668.2 of this group appears in releases 4.1 to 4.6
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | break | 6 | 7 | 8 | 1-8 | 2-7 | 5-6 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_9JA9_002 | 9JA9 | 0.3447 | 1 | A | Theophylline aptamer | C | 20 | C | 21 | C | 22 | U | 23 | G | 25 | * | C | 9 | A | 10 | G | 11 | cWW | ncWW | cWW | |
| 2 | IL_6E8U_001 | 6E8U | 0.0000 | 1 | Isolated non-canonical cWW pair | B | RNA (37-MER) | A | 6 | A | 7 | G | 8 | G | 9 | A | 11 | * | U | 30 | G | 31 | U | 32 | cWW | cWW | cWW |
| 3 | IL_3E5C_001 | 3E5C | 0.7631 | 1 | Isolated non-canonical cWW pair | A | SAM-III riboswitch aptamer (SMK box) | A | 27 | G | 28 | A | 29 | G | 31 | C | 32 | * | G | 12 | A | 13 | U | 14 | cWW | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CCCUUG*CAG | 1 |
| AAGGAA*UGU | 1 |
| AGAUGC*GAU | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| CCUU*A | 1 |
| AGGA*G | 1 |
| GAUG*A | 1 |
- Annotations
-
- Isolated non-canonical cWW pair (2)
- (1)
- Basepair signature
- cWW-cWW-L-cWW-L
- Heat map statistics
- Min 0.34 | Avg 0.43 | Max 0.84
Coloring options: