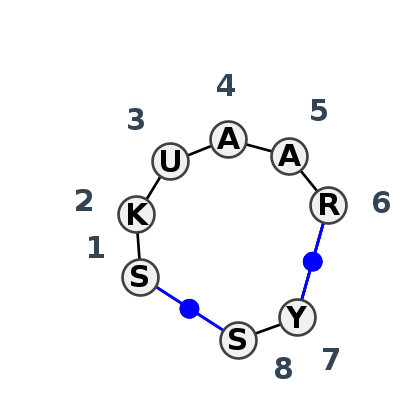
Motif IL_20847.2 Version IL_20847.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | break | 7 | 8 | 1-8 | 6-7 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_4LFB_022 | 4LFB | 0.1075 | 1 | A | SSU rRNA | C | 569 | G | 570 | U | 571 | A | 573 | A | 574 | G | 575 | * | C | 880 | G | 881 | cWW | cWW |
| 2 | IL_8B0X_025 | 8B0X | 0.0978 | 1 | A | SSU rRNA | C | 569 | G | 570 | U | 571 | A | 573 | A | 574 | G | 575 | * | C | 880 | G | 881 | cWW | cWW |
| 3 | IL_8GLP_226 | 8GLP | 0.0527 | 1 | S2 | SSU rRNA | G | 665 | U | 666 | U | 667 | A | 669 | A | 670 | A | 671 | * | U | 1161 | C | 1162 | cWW | cWW |
| 4 | IL_9H3G_189 | 9H3G | 0.0000 | 1 | h1 | SSU rRNA | G | 618 | U | 619 | U | 620 | A | 622 | A | 623 | A | 624 | * | U | 1105 | C | 1106 | cWW | cWW |
| 5 | IL_8CRE_424 | 8CRE | 0.0785 | 1 | CM | SSU rRNA | G | 614 | U | 615 | U | 616 | A | 618 | A | 619 | A | 620 | * | U | 1089 | C | 1090 | cWW | cWW |
| 6 | IL_8P9A_405 | 8P9A | 0.1117 | 1 | sR | SSU rRNA | G | 616 | U | 617 | U | 618 | A | 620 | A | 621 | A | 622 | * | U | 1104 | C | 1105 | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CGUAAAG*CG | 2 |
| GUUAAAA*UC | 2 |
| GUUAAA*UC | 1 |
| GUU(A2M)AAA*UC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GUAAA* | 2 |
| UU(A2M)AA* | 2 |
| UUAAA* | 2 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (6)
- Basepair signature
- cWW-L-cWW-L-L-R
- Heat map statistics
- Min 0.05 | Avg 0.09 | Max 0.15
Coloring options: