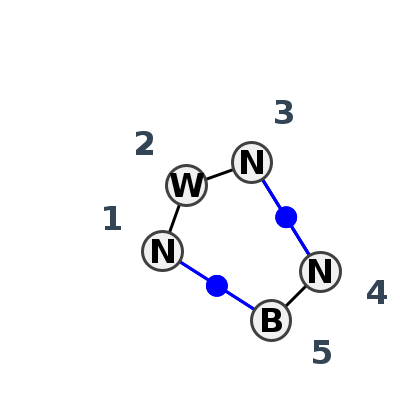
Motif IL_68541.1 Version IL_68541.1 of this group appears in releases 3.90 to 3.90
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | break | 4 | 5 | 1-5 | 3-4 | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_5J7L_267 | 5J7L | 0.8329 | 1 | 90 degree turn | DA | LSU rRNA | A | 844 | A | 845||A | U | 847||A | * | A | 933 | U | 934 | cWW | cWW |
| 2 | IL_7A0S_024 | 7A0S | 0.6312 | 0 | 90 degree turn | X | LSU rRNA | U | 942 | U | 943 | A | 944 | * | U | 860 | G | 861 | cWW | cWW |
| 3 | IL_5IBB_340 | 5IBB | 0.3361 | 0 | Single base bend | 1L | tRNA | G | 63 | U | 64 | C | 65 | * | G | 50 | C | 51 | cWW | cWW |
| 4 | IL_5J7L_266 | 5J7L | 0.1373 | 0 | Single stack bend | DA | LSU rRNA | G | 940 | A | 941 | G | 942 | * | C | 837 | C | 838 | cWW | ncWW |
| 5 | IL_8VTW_023 | 8VTW | 0.1542 | 0 | Single stack bend (H) | 1A | LSU rRNA | G | 940 | A | 941 | G | 942 | * | C | 837 | C | 838 | cWW | cWW |
| 6 | IL_4WF9_028 | 4WF9 | 0.1447 | 0 | Single stack bend | X | LSU rRNA | G | 984 | A | 985 | G | 986 | * | C | 882 | C | 883 | cWW | cWW |
| 7 | IL_7A0S_022 | 7A0S | 0.0000 | 0 | Single stack bend | X | LSU rRNA | G | 951 | A | 952 | G | 953 | * | C | 850 | C | 851 | cWW | cWW |
| 8 | IL_3MOJ_004 | 3MOJ | 0.2637 | 0 | Single stack bend | A | LSU rRNA | C | 2540 | A | 2541 | A | 2542 | * | U | 2522 | G | 2523 | cWW | cWW |
| 9 | IL_6FZ0_001 | 6FZ0 | 0.4493 | 0 | Single stack bend with SAM intercalation | A | SAM-V riboswitch | C | 46 | U | 47 | A | 48 | * | U | 20 | G | 21 | cWW | cWW |
| 10 | IL_2QWY_008 | 2QWY | 0.4724 | 0 | Single stack bend with SAM intercalation | C | SAM-II riboswitch | C | 43 | U | 44 | A | 45 | * | U | 21 | G | 22 | cWW | cWW |
| 11 | IL_3WBM_004 | 3WBM | 0.8132 | 0 | Other IL | Y+X | RNA (25-MER) | G | 2 | U | 3 | A | 4 | * | U | 23 | C | 24 | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GAG*CC | 4 |
| CUA*UG | 2 |
| AU*AAUU | 1 |
| UUA*UG | 1 |
| GUC*GC | 1 |
| CAA*UG | 1 |
| GUA*UC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| U* | 5 |
| A* | 5 |
| *AU | 1 |
Release history
| Release | 3.90 |
|---|---|
| Date | 2024-11-06 |
| Status | New id, > 2 parents |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- Single stack bend (4)
- Single stack bend with SAM intercalation (2)
- 90 degree turn (2)
- Other IL (1)
- Single base bend (1)
- Single stack bend (H) (1)
- Basepair signature
- cWW-L-cWW
- Heat map statistics
- Min 0.08 | Avg 0.50 | Max 0.90
Coloring options: