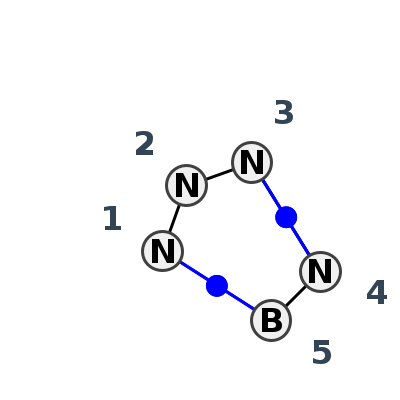
Motif IL_77710.2 Version IL_77710.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | break | 4 | 5 | 1-5 | 3-4 | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_3DD2_003 | 3DD2 | 0.4773 | 1 | Major groove intercalation | B | RNA (26-MER) | G | 16 | A | 18 | CFL | 19 | * | G | 10 | CFL | 11 | cWW | cWW |
| 2 | IL_4WF9_106 | 4WF9 | 0.3279 | 0 | Major groove platform | X | LSU rRNA | U | 2898 | A | 2899 | C | 2900 | * | G | 2858 | G | 2859 | cWW | cWW |
| 3 | IL_9E6Q_101 | 9E6Q | 0.3402 | 0 | 1 | LSU rRNA | U | 3005 | C | 3006 | C | 3007 | * | G | 2957 | G | 2958 | cWW | cWW | |
| 4 | IL_8B0X_168 | 8B0X | 0.3360 | 0 | Major groove intercalation (H) | a | LSU rRNA | U | 2882 | A | 2883 | C | 2884 | * | G | 2842 | G | 2843 | cWW | cWW |
| 5 | IL_5WT1_001 | 5WT1 | 0.3134 | 0 | Stack outside cWW | C | tRNA | G | 43 | G | 44 | U | 45 | * | A | 26 | C | 27 | cWW | cWW |
| 6 | IL_7VTI_003 | 7VTI | 0.5286 | 1 | Major groove intercalation | B | crRNA | U | 27 | A | 29 | U | 30 | * | A | 9 | G | 10 | cWW | cWW |
| 7 | IL_7A0S_067 | 7A0S | 0.3567 | 1 | Major groove intercalation | X | LSU rRNA | U | 1881 | G | 1882 | A | 1884 | * | U | 1833 | G | 1834 | cWW | cWW |
| 8 | IL_6JQ6_004 | 6JQ6 | 0.2580 | 0 | Stack outside cWW | U | Hatchet ribozyme | U | 62 | G | 63 | C | 64 | * | G | 31 | G | 32 | cWW | cWW |
| 9 | IL_7MLW_005 | 7MLW | 0.2495 | 0 | Stack outside cWW | F | Guanidine-I riboswitch | U | 43 | G | 44 | C | 45 | * | G | 17 | G | 18 | cWW | cWW |
| 10 | IL_9DFE_072 | 9DFE | 0.2417 | 1 | Stack outside cWW (H) | 1A | LSU rRNA | U | 1898 | G | 1899 | A | 1901 | * | U | 1841 | G | 1842 | cWW | cWW |
| 11 | IL_6CF2_002 | 6CF2 | 0.1617 | 1 | Major groove intercalation | G | RNA (35-MER) | C | 22 | U | 23 | G | 25 | * | C | 10 | G | 11 | cWW | cWW |
| 12 | IL_6DLR_004 | 6DLR | 0.1475 | 1 | Multiple bulged bases | A | Guanidine-I riboswitch | G | 36 | G | 37 | C | 39 | * | G | 22 | C | 23 | cWW | cWW |
| 13 | IL_6XH2_001 | 6XH2 | 0.1668 | 2 | Major groove intercalation | D | TRANS-ACTIVATION RESPONSE ELEMENT | A | 22 | U | 23 | G | 26 | * | C | 39 | U | 40 | cWW | cWW |
| 14 | IL_1XOK_002 | 1XOK | 0.1500 | 1 | Major groove intercalation | B+A | alfalfa mosaic virus RNA 3' UTR | G | 877 | A | 878 | G | 880 | * | C | 868 | C | 869 | cWW | cWW |
| 15 | IL_1XOK_001 | 1XOK | 0.1491 | 1 | Major groove intercalation | A | alfalfa mosaic virus RNA 3' UTR | A | 864 | A | 865 | G | 867 | * | C | 846 | U | 847 | cWW | cWW |
| 16 | IL_5LYS_007 | 5LYS | 0.1175 | 1 | Major groove intercalation | B | 7SK RNA | A | 39 | U | 40 | G | 42 | * | C | 67 | U | 68 | cWW | cWW |
| 17 | IL_5T83_004 | 5T83 | 0.0000 | 1 | Multiple bulged bases | A | Guanidine-I riboswitch | G | 33 | G | 34 | C | 36 | * | G | 22 | C | 23 | cWW | cWW |
| 18 | IL_4KZD_006 | 4KZD | 0.1849 | 1 | Stack outside cWW | R | RNA (84-MER) | G | 49 | U | 50 | G | 52 | * | C | 30 | C | 31 | cWW | cWW |
| 19 | IL_5LYS_006 | 5LYS | 0.1852 | 1 | Stack outside cWW | B | 7SK RNA | G | 70 | C | 71 | G | 73 | * | C | 36 | C | 37 | cWW | cWW |
| 20 | IL_8K7W_004 | 8K7W | 0.1640 | 1 | A | RNA (49-MER) | G | 30 | U | 31 | G | 33 | * | C | 14 | C | 15 | cWW | cWW | |
| 21 | IL_8K85_004 | 8K85 | 0.2034 | 1 | B+A | RNA (56-MER) | G | 36 | U | 37 | G | 39 | * | C | 15 | C | 16 | cWW | cWW | |
| 22 | IL_1U9S_008 | 1U9S | 0.2488 | 0 | Stack outside cWW | A | RNase P class A | G | 217 | G | 218 | C | 219 | * | G | 195 | C | 196 | cWW | cWW |
| 23 | IL_7L0Z_004 | 7L0Z | 0.2181 | 1 | Stack outside cWW | G | RNA (69-MER) | G | 41 | U | 42 | G | 44 | * | C | 23 | C | 24 | cWW | cWW |
| 24 | IL_8K85_009 | 8K85 | 0.2521 | 1 | A+B | RNA (56-MER) | G | 36 | U | 37 | G | 39 | * | C | 15 | C | 16 | cWW | cWW | |
| 25 | IL_9H3G_019 | 9H3G | 0.2992 | 1 | A | LSU rRNA | C | 433 | G | 434 | C | 436 | * | G | 633 | G | 634 | cWW | cWW | |
| 26 | IL_1NTA_001 | 1NTA | 0.3513 | 0 | Stack outside cWW | A+B | RNA (22-mer) + RNA (18-mer) | C | 5 | G | 6 | C | 7 | * | G | 114 | G | 115 | cWW | cWW |
| 27 | IL_7OAX_001 | 7OAX | 0.4151 | 0 | Stack outside cWW | A | Chili RNA Aptamer | U | 8 | G | 9 | G | 10 | * | C | 44 | G | 45 | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GUUG*CC | 4 |
| UAC*GG | 2 |
| UGAA*UG | 2 |
| UGC*GG | 2 |
| G(UFT)A(CFL)*G(CFL) | 1 |
| UCC*GG | 1 |
| GGU*AC | 1 |
| UGAU*AG | 1 |
| CUGG*CG | 1 |
| GGAC*GC | 1 |
| AUCUG*CU | 1 |
| GAUG*CC | 1 |
| AAUG*CU | 1 |
| AUUG*CU | 1 |
| GGUC*GC | 1 |
| GCUG*CC | 1 |
| GUCG*CC | 1 |
| GGC*GC | 1 |
| CGGC*GG | 1 |
| CGC*GG | 1 |
| UGG*CG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| G* | 6 |
| UU* | 5 |
| GA* | 4 |
| A* | 2 |
| AU* | 2 |
| (UFT)A* | 1 |
| C* | 1 |
| UG* | 1 |
| UCU* | 1 |
| GU* | 1 |
| CU* | 1 |
| UC* | 1 |
| GG* | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- Stack outside cWW (9)
- Major groove intercalation (8)
- (5)
- Multiple bulged bases (2)
- Major groove platform (1)
- Major groove intercalation (H) (1)
- Stack outside cWW (H) (1)
- Basepair signature
- cWW-L-cWW
- Heat map statistics
- Min 0.08 | Avg 0.34 | Max 0.92
Coloring options: