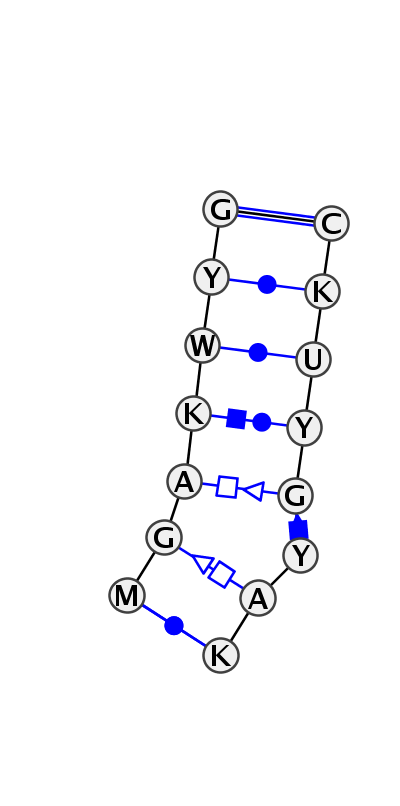
Motif IL_90248.1 Version IL_90248.1 of this group appears in releases 2.0 to 2.0
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | break | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 1-15 | 2-14 | 3-12 | 4-11 | 5-9 | 5-10 | 6-9 | 7-8 | 12-13 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_3J79_025 | 3J79 | 0.0000 | 0 | A | LSU rRNA | A | 827 | G | 828 | A | 829 | U | 830 | U | 831 | U | 832 | G | 833 | * | C | 861 | U | 862 | U | 863 | U | 864 | G | 865 | C | 866 | A | 867 | U | 868 | cWW | tSH | ntHS | cHW | cWw | ncwW | cWW | ||
| 2 | IL_3J7P_203 | 3J7P | 0.4171 | 1 | S2 | SSU rRNA | C | 1047 | G | 1048 | A | 1050 | G | 1051 | A | 1052 | C | 1053 | G | 1054 | * | C | 1064 | G | 1065 | U | 1066 | C | 1067 | G | 1068 | U | 1069 | A | 1070 | G | 1071 | cWW | ntSH | ncWW | cWW | cSH |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AGAUUUG*CUUUGCAU | 1 |
| CGAAGACG*CGUCGUAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAUUU*UUUGCA | 1 |
| GAAGAC*GUCGUA | 1 |
Release history
| Release | 2.0 |
|---|---|
| Date | 2017-04-24 |
| Status | New id, no parents |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (2)
- Basepair signature
- cWW-tSH-cSH-tHS-cHW-cWW-cWW-cWW
- Heat map statistics
- Min 0.42 | Avg 0.21 | Max 0.42
Coloring options: