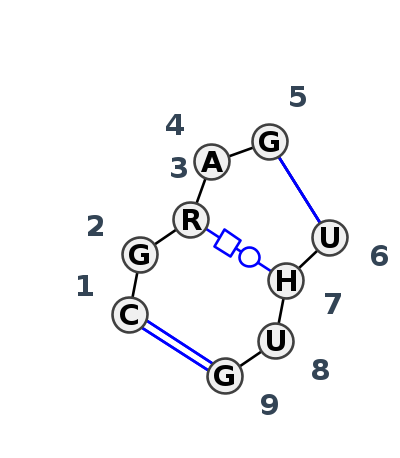
Motif IL_98200.2 Version IL_98200.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | break | 6 | 7 | 8 | 9 | 1-9 | 2-8 | 3-7 | 5-6 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_9H3G_131 | 9H3G | 0.1511 | 2 | A | LSU rRNA | C | 3082 | G | 3083 | A | 3084 | A | 3085 | G | 3086 | * | U | 3052 | C | 3053 | U | 3055 | G | 3057 | cWW | tHW | cWW | ||
| 2 | IL_9AXU_111 | 9AXU | 0.0000 | 2 | 2 | LSU rRNA | C | 3180 | G | 3181 | G | 3182 | A | 3183 | G | 3184 | * | U | 3150 | A | 3151 | U | 3153 | G | 3155 | cWW | cWW | |||
| 3 | IL_8P9A_346 | 8P9A | 0.1246 | 2 | UAA/GAN related (H) | AR | LSU rRNA | C | 3084 | G | 3085 | A | 3086 | A | 3087 | G | 3088 | * | U | 3054 | U | 3055 | U | 3057 | G | 3059 | cWW | ntHW | cWW | |
| 4 | IL_8OI5_118 | 8OI5 | 0.1593 | 2 | UAA/GAN related (H) | 1 | LSU rRNA | C | 3056 | G | 3057 | A | 3058 | A | 3059 | G | 3060 | * | U | 3026 | U | 3027 | U | 3029 | G | 3031 | cWW | tSW | tHW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CGAAG*UUUUUG | 2 |
| CGAAG*UCAUCG | 1 |
| CGGAG*UAUUUG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAA*UUUU | 2 |
| GAA*CAUC | 1 |
| GGA*AUUU | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- UAA/GAN related (H) (2)
- (2)
- Basepair signature
- cWW-L-R-tHW-L-cWW
- Heat map statistics
- Min 0.12 | Avg 0.11 | Max 0.19
Coloring options: