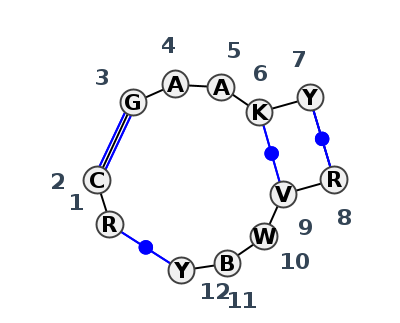
Motif J3_01004.2 Version J3_01004.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | break | 3 | 4 | 5 | 6 | 7 | break | 8 | 9 | 10 | 11 | 12 | 1-12 | 2-3 | 3-4 | 6-9 | 7-8 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | J3_8GLP_039 | 8GLP | 0.1709 | 2 | L8 | 5.8S rRNA | A | 44 | C | 45 | * | G | 58 | A | 59 | A | 61 | U | 63 | U | 64 | * | A | 95 | C | 96 | A | 97 | C | 98 | U | 99 | cWW | cWW | cWW | ||
| 2 | J3_9H3G_004 | 9H3G | 0.1473 | 2 | 3 | 5.8S rRNA | A | 48 | C | 49 | * | G | 62 | A | 63 | A | 65 | U | 67 | U | 68 | * | A | 100 | G | 101 | U | 102 | C | 103 | U | 104 | cWW | cWW | ncSs | cWW | |
| 3 | J3_8P9A_067 | 8P9A | 0.1346 | 2 | AT | 5.8S rRNA | A | 44 | C | 45 | * | G | 58 | A | 59 | A | 61 | G | 63 | U | 64 | * | A | 96 | A | 97 | U | 98 | C | 99 | U | 100 | cWW | cWW | ncSs | cWW | cWW |
| 4 | J3_8OI5_026 | 8OI5 | 0.1415 | 2 | 4 | 5.8S rRNA | A | 44 | C | 45 | * | G | 58 | A | 59 | A | 61 | G | 63 | U | 64 | * | A | 96 | A | 97 | U | 98 | C | 99 | U | 100 | cWW | cWW | ncSs | cWW | cWW |
| 5 | J3_9AXU_027 | 9AXU | 0.1318 | 2 | 4 | 5.8S rRNA | A | 52 | C | 53 | * | G | 66 | A | 67 | A | 69 | G | 71 | U | 72 | * | A | 104 | A | 105 | U | 106 | C | 107 | U | 108 | cWW | cWW | ncSs | cWW | cWW |
| 6 | J3_8B0X_015 | 8B0X | 0.0000 | 2 | a | LSU rRNA | A | 56 | C | 57 | * | G | 70 | A | 71 | A | 73 | G | 75 | C | 76 | * | G | 110 | A | 111 | U | 112 | U | 113 | U | 114 | cWW | cWW | ncSs | cWW | cWW |
| 7 | J3_9E6Q_004 | 9E6Q | 0.0572 | 2 | 1 | LSU rRNA | G | 48 | C | 49 | * | G | 62 | A | 63 | A | 65 | G | 67 | C | 68 | * | G | 101 | A | 102 | U | 103 | C | 104 | C | 105 | cWW | cWW | cWW | cWW | |
| 8 | J3_4V9F_014 | 4V9F | 0.0547 | 2 | 0 | LSU rRNA | A | 52 | C | 53 | * | G | 66 | A | 67 | A | 69 | G | 71 | C | 72 | * | G | 105 | A | 106 | U | 107 | U | 108 | U | 109 | cWW | cWW | ncSs | cWW | cWW |
| 9 | J3_9DFE_004 | 9DFE | 0.1288 | 2 | 1A | LSU rRNA | A | 56 | C | 57 | * | G | 70 | A | 71 | A | 73 | G | 75 | C | 76 | * | G | 110 | A | 111 | U | 112 | G | 113 | U | 114 | cWW | cWW | ncSs | cWW | cWW |
| 10 | J3_7A0S_002 | 7A0S | 0.1570 | 2 | X | LSU rRNA | A | 55 | C | 56 | * | G | 69 | A | 70 | A | 72 | G | 74 | C | 75 | * | G | 108 | A | 109 | U | 110 | G | 111 | U | 112 | cWW | cWW | ncWW | cWW | |
| 11 | J3_4WF9_013 | 4WF9 | 0.1743 | 2 | X | LSU rRNA | A | 56 | C | 57 | * | G | 70 | A | 71 | A | 73 | G | 75 | C | 76 | * | G | 109 | A | 110 | U | 111 | U | 112 | U | 113 | cWW | cWW | cSH | cWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AC*GAUACGU*AAUCU | 3 |
| AC*GAUAAGC*GAUUU | 2 |
| AC*GAGAAUU*ACACU | 1 |
| AC*GAUACUU*AGUCU | 1 |
| GC*GAUAAGC*GAUCC | 1 |
| AC*GAUAAGC*GAUGU | 1 |
| AC*GAAAAGC*GAUGU | 1 |
| AC*GAUAUGC*GAUUU | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| *AUACG*AUC | 3 |
| *AUAAG*AUU | 2 |
| *AGAAU*CAC | 1 |
| *AUACU*GUC | 1 |
| *AUAAG*AUC | 1 |
| *AUAAG*AUG | 1 |
| *AAAAG*AUG | 1 |
| *AUAUG*AUU | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (11)
- Basepair signature
- cWW-cWW-F-F-F-cWW-F-cWW
- Heat map statistics
- Min 0.05 | Avg 0.14 | Max 0.25
Coloring options: