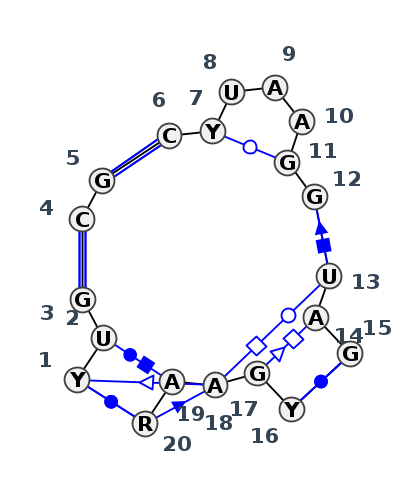
Motif J4_42306.2 Version J4_42306.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | break | 4 | 5 | break | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | break | 16 | 17 | 18 | 19 | 20 | 1-18 | 1-20 | 2-19 | 3-4 | 5-6 | 7-11 | 8-9 | 8-10 | 12-13 | 13-18 | 14-17 | 15-16 | 18-20 | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | J4_8B0X_012 | 8B0X | 0.3765 | 1 | a | LSU rRNA | C | 1835 | U | 1836 | 2MG | 1837 | * | C | 1909 | G | 1910 | * | C | 1928 | C | 1929 | U | 1930 | A | 1931 | A | 1932 | G | 1933 | G | 1934 | U | 1935 | A | 1936 | G | 1937 | * | C | 1971 | G | 1972 | A | 1973 | A | 1974 | G | 1976 | tsS | cWW | cWH | cWW | cWW | tWW | cSH | ntWH | tHS | cWW | cSs | ||
| 2 | J4_9DFE_008 | 9DFE | 0.3723 | 1 | 1A | LSU rRNA | U | 1833 | U | 1834 | G | 1835 | * | C | 1905 | G | 1906 | * | C | 1924 | C | 1925 | U | 1926 | A | 1927 | A | 1928 | G | 1929 | G | 1930 | U | 1931 | A | 1932 | G | 1933 | * | C | 1967 | G | 1968 | A | 1969 | A | 1970 | A | 1972 | tsS | cWW | cWH | cWW | cWW | tWW | cSH | tWH | tHS | cWW | |||
| 3 | J4_9H3G_008 | 9H3G | 0.0000 | 1 | A | LSU rRNA | C | 2190 | U | 2191 | G | 2192 | * | C | 2246 | G | 2247 | * | C | 2265 | U | 2266 | U | 2267 | A | 2268 | A | 2269 | G | 2270 | G | 2271 | U | 2272 | A | 2273 | G | 2274 | * | U | 2308 | G | 2309 | A | 2310 | A | 2311 | G | 2313 | tsS | cWW | cWH | cWW | cWW | tWW | ncSH | cSH | tWH | tHS | cWW | cSs | |
| 4 | J4_4V9F_008 | 4V9F | 0.1086 | 1 | 0 | LSU rRNA | C | 1889 | U | 1890 | G | 1891 | * | C | 1946 | G | 1947 | * | C | 1965 | U | 1966 | U | 1967 | A | 1968 | A | 1969 | G | 1970 | G | 1971 | U | 1972 | A | 1973 | G | 1974 | * | U | 2008 | G | 2009 | A | 2010 | A | 2011 | G | 2013 | tsS | cWW | cWH | cWW | cWW | tWW | cSH | tWH | tHS | cWW | cSs | ||
| 5 | J4_8P9A_018 | 8P9A | 0.1154 | 1 | AR | LSU rRNA | C | 2192 | U | 2193 | G | 2194 | * | C | 2248 | G | 2249 | * | C | 2267 | U | 2268 | U | 2269 | A | 2270 | A | 2271 | G | 2272 | G | 2273 | U | 2274 | A | 2275 | G | 2276 | * | U | 2310 | G | 2311 | A | 2312 | A | 2313 | G | 2315 | tsS | cWW | cWH | cWW | cWW | ntWW | ncSH | cSH | tWH | tHS | cWW | cSs | |
| 6 | J4_9E6Q_008 | 9E6Q | 0.1766 | 1 | 1 | LSU rRNA | C | 1969 | U | 1970 | OMG | 1971 | * | C | 2026 | G | 2027 | * | C | 2045 | U | 2046 | U | 2047 | A | 2048 | A | 2049 | G | 2050 | G | 2051 | U | 2052 | A | 2053 | G | 2054 | * | OMU | 2088 | G | 2089 | A | 2090 | A | 2091 | G | 2093 | tsS | cWW | cWH | cWW | cWW | tWW | cSH | tWH | tHS | cWW | cSs |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CUG*CG*CUUAAGGUAG*UGAAUG | 2 |
| CU2MG*CG*CCUAAGGUAG*CGAAUG | 1 |
| UUG*CG*CCUAAGGUAG*CGAAAA | 1 |
| CUG*CG*CUUAAGGUAG*UGAA(PSU)G | 1 |
| CUOMG*CG*CUUAAGGUAG*OMUGAAUG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| U**UUAAGGUA*GAAU | 3 |
| U**CUAAGGUA*GAAU | 1 |
| U**CUAAGGUA*GAAA | 1 |
| U**UUAAGGUA*GAA(PSU) | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (6)
- Basepair signature
- cWW-tSS-cSS-cWH-cWW-tWH-tHS-cWW-cWW-tWW-F-cSH-F-F
- Heat map statistics
- Min 0.07 | Avg 0.24 | Max 0.47
Coloring options: