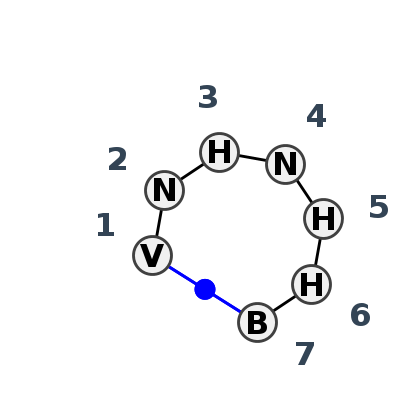
Motif HL_10453.2 Version HL_10453.2 of this group appears in releases 3.95 to 3.95
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 1-7 | 2-6 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_4V9F_049 | 4V9F | 0.5835 | 1 | GNRA related | 0 | LSU rRNA | C | 1916 | G | 1917 | A | 1919 | C | 1920 | A | 1921 | A | 1922 | G | 1923 | cWW | ntSH |
| 2 | HL_7A0S_029 | 7A0S | 0.4866 | 1 | GNRA related | X | LSU rRNA | C | 1103 | G | 1104 | A | 1106 | A | 1107 | U | 1108 | A | 1109 | G | 1110 | cWW | ntSH |
| 3 | HL_3OXE_001 | 3OXE | 0.5106 | 1 | Anticodon loop related | A | Glycine riboswitch | G | 17 | U | 19 | U | 20 | A | 21 | A | 22 | U | 23 | C | 24 | cWW | |
| 4 | HL_6UFH_001 | 6UFH | 0.4058 | 1 | Anticodon loop related | A | RNA (167-MER) | G | 13 | C | 14 | A | 16 | U | 17 | C | 18 | A | 19 | C | 20 | cWW | |
| 5 | HL_8H1B_001 | 8H1B | 0.2781 | 2 | tRNA anticodon loop | C | RNA (17-mer) | A | 31 | C | 32 | U | 35 | G | 36 | A | 37 | C | 38 | U | 39 | cWW | ncWB |
| 6 | HL_8H0S_002 | 8H0S | 0.2217 | 2 | tRNA anticodon loop | Y | RNA (17-mer) | A | 31 | C | 32 | U | 35 | G | 36 | A | 37 | C | 38 | U | 39 | cWW | ncWB |
| 7 | HL_6MPI_033 | 6MPI | 0.1038 | 2 | tRNA anticodon loop | X | tRNA ASL Escherichia coli Arg1 | G | 31 | RSP | 32 | C | 35 | G | 36 | A | 37 | A | 38 | C | 39 | cWW | |
| 8 | HL_6MKN_034 | 6MKN | 0.0000 | 2 | tRNA anticodon loop | X | tRNA ASL Escherichia coli Arg2 | G | 31 | C | 32 | C | 35 | G | 36 | A | 37 | A | 38 | C | 39 | cWW | |
| 9 | HL_5U3G_003 | 5U3G | 0.7082 | 2 | B | Guanidine-I riboswitch | G | 68 | A | 69 | A | 71 | A | 72 | A | 73 | A | 74 | C | 76 | cWW | tHH |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| ACUUUGACU | 2 |
| CGUACAAG | 1 |
| CGUAAUAG | 1 |
| GUUUAAUC | 1 |
| GCGAUCAC | 1 |
| G(RSP)U(I)CGAAC | 1 |
| GCU(I)CGAAC | 1 |
| GAAAAAAGC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| CUUUGAC | 2 |
| GUACAA | 1 |
| GUAAUA | 1 |
| UUUAAU | 1 |
| CGAUCA | 1 |
| (RSP)U(I)CGAA | 1 |
| CU(I)CGAA | 1 |
| AAAAAAG | 1 |
Release history
| Release | 3.95 |
|---|---|
| Date | 2025-03-26 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- tRNA anticodon loop (4)
- Anticodon loop related (2)
- GNRA related (2)
- (1)
- Basepair signature
- cWW-F-F-F-F-F
- Heat map statistics
- Min 0.10 | Avg 0.49 | Max 0.97
Coloring options: