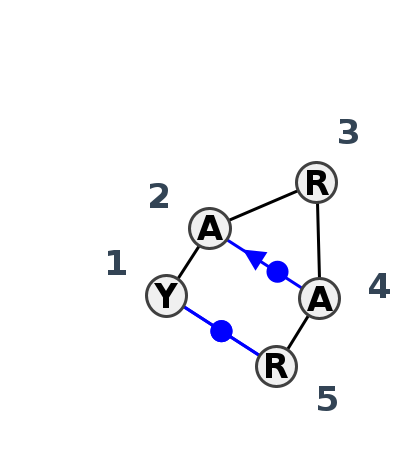
Motif HL_19413.2 Version HL_19413.2 of this group appears in releases 4.1 to 4.6
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 1-5 | 2-4 | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_9F37_001 | 9F37 | 0.0883 | 1 | V | 5' end of the vRNA promoter | U | 3 | A | 4 | G | 5 | A | 7 | A | 8 | cWW | ncSW |
| 2 | HL_7Z42_002 | 7Z42 | 0.0782 | 1 | V | RNA (13-mer) | U | 3 | A | 4 | G | 5 | A | 7 | A | 8 | cWW | cSW |
| 3 | HL_5FMZ_001 | 5FMZ | 0.0717 | 1 | H | RNA (12-mer) | U | 3 | A | 4 | G | 5 | A | 7 | A | 8 | cWW | |
| 4 | HL_4WRT_001 | 4WRT | 0.0000 | 1 | V | Influenza virus polymerase vRNA promoter 5' end | U | 3 | A | 4 | G | 5 | A | 7 | A | 8 | cWW | ncSW |
| 5 | HL_4WSB_001 | 4WSB | 0.0724 | 1 | V | Influenza A polymerase vRNA promoter 5' end | U | 3 | A | 4 | G | 5 | A | 7 | A | 8 | cWW | |
| 6 | HL_7Z4O_002 | 7Z4O | 0.1615 | 1 | VVV | RNA (12-mer) | U | 3 | A | 4 | G | 5 | A | 7 | A | 8 | cWW | |
| 7 | HL_7EL1_002 | 7EL1 | 0.2635 | 2 | B | RNA (73-MER) | C | 59 | A | 60 | A | 61 | A | 63 | G | 65 | cWW | cSW |
| 8 | HL_7ENI_002 | 7ENI | 0.2623 | 2 | B | sgRNA (74-MER) | C | 59 | A | 60 | A | 61 | A | 63 | G | 65 | cWW | cSW |
| 9 | HL_9E6Q_007 | 9E6Q | 0.6592 | 1 | 1 | LSU rRNA | C | 206 | A | 207 | A | 208 | A | 210 | G | 211 | cWW | cSW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UAGUAA | 6 |
| CAAAAUG | 2 |
| CAACAG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AGUA | 6 |
| AAAAU | 2 |
| AACA | 1 |
- Annotations
-
- (9)
- Basepair signature
- cWW-cSW-F
- Heat map statistics
- Min 0.07 | Avg 0.25 | Max 0.68
Coloring options: