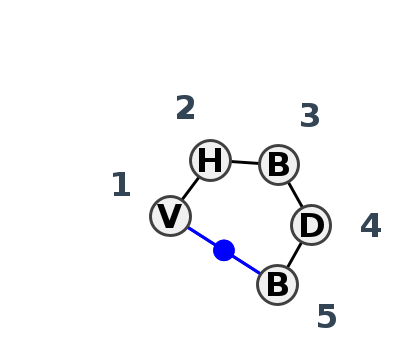
Motif HL_45127.1 Version HL_45127.1 of this group appears in releases 3.91 to 3.94
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 1-5 | 2-4 | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_8UO6_003 | 8UO6 | 0.9234 | 2 | A | HIV-1 Rev Response Element Stem-Loop II with tRNA scaffold | A | 77 | A | 78 | U | 79 | U | 82 | U | 83 | cWW | ||
| 2 | HL_8P9A_179 | 8P9A | 0.2284 | 2 | AR | LSU rRNA | C | 3193 | C | 3194 | U | 3196 | G | 3197 | G | 3199 | cWW | ncWW | |
| 3 | HL_5J7L_214 | 5J7L | 0.0000 | 2 | DA | LSU rRNA | C | 1727 | C | 1728 | C | 1730 | G | 1731 | G | 1733 | cWW | ncWW | |
| 4 | HL_5J7L_200 | 5J7L | 0.2469 | 2 | UNCG variation | DA | LSU rRNA | C | 2795 | U | 2796 | U | 2798 | A | 2799 | G | 2801 | cWW | ncWW |
| 5 | HL_8P9A_130 | 8P9A | 0.4623 | 1 | AR | LSU rRNA | G | 763 | U | 764 | U | 766 | U | 767 | C | 768 | cWW | cwW | |
| 6 | HL_2Y9H_005 | 2Y9H | 0.3833 | 3 | J | RNA (19-mer) | A | 10 | C | 11 | G | 12 | G | 16 | U | 17 | cWW | ncWH |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AAUUAUU | 1 |
| CCUUGUG | 1 |
| CCUCGCG | 1 |
| CUUUAAG | 1 |
| GUCUUC | 1 |
| ACGCGUGU | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AUUAU | 1 |
| CUUGU | 1 |
| CUCGC | 1 |
| UUUAA | 1 |
| UCUU | 1 |
| CGCGUG | 1 |
- Annotations
-
- (5)
- UNCG variation (1)
- Basepair signature
- cWW-F-F-F
- Heat map statistics
- Min 0.23 | Avg 0.47 | Max 0.97
Coloring options: