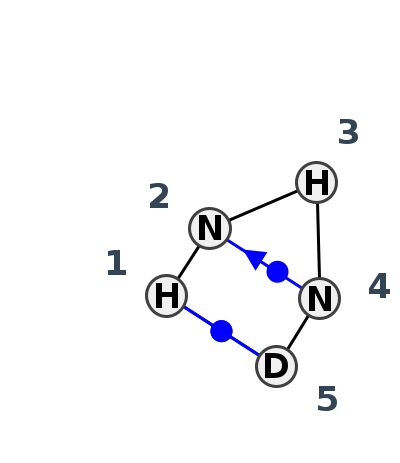
Motif HL_51020.2 Version HL_51020.2 of this group appears in releases 4.1 to 4.6
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 1-5 | 2-4 | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_4UYK_003 | 4UYK | 0.4488 | 1 | R | SRP RNA | U | 98 | G | 99 | U | 100 | A | 101 | G | 103 | cWW | ntSW | |
| 2 | HL_9DFE_006 | 9DFE | 0.2900 | 1 | 1A | LSU rRNA | A | 225 | G | 226 | A | 227 | A | 228 | U | 230 | cWW | ||
| 3 | HL_7A0S_007 | 7A0S | 0.2756 | 1 | X | LSU rRNA | A | 202 | G | 203 | A | 204 | A | 205 | U | 207 | cWW | cSW | |
| 4 | HL_4WF9_007 | 4WF9 | 0.2299 | 1 | X | LSU rRNA | A | 228 | A | 229 | A | 230 | A | 231 | U | 233 | cWW | cSW | |
| 5 | HL_9AXU_002 | 9AXU | 0.0964 | 1 | 2 | LSU rRNA | U | 69 | A | 70 | A | 71 | C | 72 | A | 74 | cWW | cSW | |
| 6 | HL_8GLP_002 | 8GLP | 0.0000 | 1 | L5 | LSU rRNA | U | 68 | A | 69 | A | 70 | C | 71 | A | 73 | cWW | cSW | |
| 7 | HL_8OI5_002 | 8OI5 | 0.0948 | 1 | 1 | LSU rRNA | C | 68 | A | 69 | A | 70 | C | 71 | G | 73 | cWW | cSW | |
| 8 | HL_8P9A_114 | 8P9A | 0.1149 | 1 | AR | LSU rRNA | C | 69 | A | 70 | A | 71 | C | 72 | G | 74 | cWW | cSW | |
| 9 | HL_8P9A_193 | 8P9A | 0.3373 | 1 | sR | SSU rRNA | C | 190 | C | 191 | U | 192 | U | 193 | G | 195 | cWW | ||
| 10 | HL_2NUE_001 | 2NUE | 0.4448 | 1 | UNCG variation | C | 46-MER | C | 21 | G | 22 | A | 24 | A | 25 | G | 26 | cWW | ncSW |
| 11 | HL_7A0S_023 | 7A0S | 0.5076 | 2 | UNCG variation | X | LSU rRNA | U | 792 | G | 793 | A | 795 | A | 796 | G | 798 | cWW | cSW |
| 12 | HL_6VMY_004 | 6VMY | 0.5057 | 2 | UNCG variation | A | Cobalamin riboswitch | C | 287 | C | 288 | A | 290 | G | 291 | G | 293 | cWW | ncWH |
| 13 | HL_8S95_003 | 8S95 | 0.5837 | 2 | UNCG variation | C | Enterovirus 5' cloverleaf cis-acting replication element | C | 67 | C | 68 | C | 70 | G | 71 | G | 73 | cWW | ncWH |
| 14 | HL_8P9A_194 | 8P9A | 0.7007 | 1 | UNCG variation | sR | SSU rRNA | C | 230 | U | 231 | C | 233 | G | 234 | G | 235 | ncWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UAACCA | 2 |
| UGUAAG | 1 |
| AGAAAU | 1 |
| AGAAUU | 1 |
| AAAAUU | 1 |
| CAACAG | 1 |
| CAACCG | 1 |
| CCUUUG | 1 |
| CGCAAG | 1 |
| UGAAAAG | 1 |
| CCAAGCG | 1 |
| CCACGUG | 1 |
| CUUCGG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AACC | 3 |
| GUAA | 1 |
| GAAA | 1 |
| GAAU | 1 |
| AAAU | 1 |
| AACA | 1 |
| CUUU | 1 |
| GCAA | 1 |
| GAAAA | 1 |
| CAAGC | 1 |
| CACGU | 1 |
| UUCG | 1 |
- Annotations
-
- (9)
- UNCG variation (5)
- Basepair signature
- cWW-cSW-F
- Heat map statistics
- Min 0.07 | Avg 0.42 | Max 0.82
Coloring options: