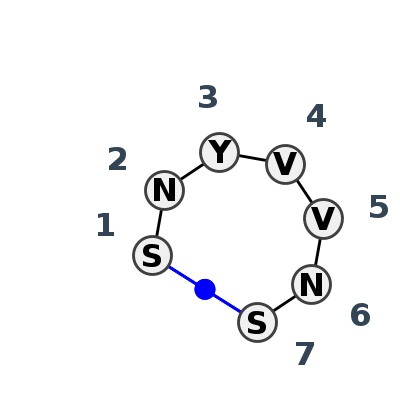
Motif HL_67772.2 Version HL_67772.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 1-7 | 2-6 | 3-5 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_8B0X_056 | 8B0X | 0.7036 | 0 | a | LSU rRNA | C | 887 | A | 888 | U | 889 | C | 890 | C | 891 | C | 892 | G | 893 | cWW | tSH | ||
| 2 | HL_9DFE_024 | 9DFE | 0.6193 | 1 | 1A | LSU rRNA | C | 884 | C | 885 | C | 886 | C | 888 | C | 889 | A | 890 | G | 892 | cWW | ncSW | ntSH | |
| 3 | HL_3IVK_005 | 3IVK | 0.5734 | 1 | T-loop related | M | class I ligase product | G | 92 | U | 93 | U | 94 | A | 95 | A | 96 | A | 98 | C | 99 | cWW | ntSH | |
| 4 | HL_7TZS_002 | 7TZS | 0.4314 | 1 | T-loop related | X | TPP riboswitch (THI element) | G | 66 | A | 67 | U | 68 | A | 69 | A | 70 | U | 71 | C | 73 | cWW | ||
| 5 | HL_3K0J_002 | 3K0J | 0.3946 | 1 | T-loop related | E | TPP riboswitch (THI element) | G | 66 | A | 67 | U | 68 | A | 69 | A | 70 | U | 71 | C | 73 | cWW | ||
| 6 | HL_3D2V_002 | 3D2V | 0.4295 | 1 | T-loop related | A | TPP riboswitch (THI element) | G | 54 | G | 55 | U | 56 | A | 57 | A | 58 | U | 59 | C | 61 | cWW | ||
| 7 | HL_5CCB_003 | 5CCB | 0.4375 | 2 | T-loop related | N | tRNA | G | 53 | U | 54 | U | 55 | C | 56 | A | 58||A | G | 59 | C | 61 | cWW | ||
| 8 | HL_8GLP_089 | 8GLP | 0.2171 | 1 | T-loop with 1 bulged base (H) | S2 | SSU rRNA | G | 288 | G | 289 | U | 290 | G | 291 | A | 292 | C | 293 | C | 295 | cWW | ||
| 9 | HL_9H3G_083 | 9H3G | 0.1150 | 1 | h1 | SSU rRNA | OMG | 244 | A | 245 | U | 246 | G | 247 | A | 248 | U | 249 | C | 251 | cWW | |||
| 10 | HL_8CRE_196 | 8CRE | 0.0897 | 1 | T-loop with 1 bulged base | CM | SSU rRNA | G | 241 | A | 242 | U | 243 | G | 244 | A | 245 | U | 246 | C | 248 | cWW | ||
| 11 | HL_8P9A_195 | 8P9A | 0.0000 | 1 | T-loop with 1 bulged base | sR | SSU rRNA | G | 243 | A | 244 | U | 245 | G | 246 | A | 247 | U | 248 | C | 250 | cWW | ||
| 12 | HL_4V9F_039 | 4V9F | 0.3866 | 0 | Other HL | 0 | LSU rRNA | C | 1594 | G | 1595 | U | 1596 | A | 1597 | A | 1598 | U | 1599 | G | 1600 | cWW | ||
| 13 | HL_9E6Q_037 | 9E6Q | 0.5260 | 0 | Other HL (H) | 1 | LSU rRNA | C | 1638 | G | 1639 | U | 1640 | A | 1641 | A | 1642 | U | 1643 | G | 1644 | cWW | ||
| 14 | HL_8GLP_042 | 8GLP | 0.5211 | 0 | L5 | LSU rRNA | G | 2664 | U | 2665 | U | 2666 | C | 2667 | G | 2668 | C | 2669 | C | 2670 | cWW | tSH |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GAUAAUGC | 2 |
| GAUGAUUC | 2 |
| CGUAAUG | 2 |
| CAUCCCG | 1 |
| CCCACCAG | 1 |
| GUUAAAAC | 1 |
| GGUAAUGC | 1 |
| GUUCAAGUC | 1 |
| GGUGACUC | 1 |
| (OMG)AUGAUUC | 1 |
| GUUCGCC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AUGAUU | 3 |
| AUAAUG | 2 |
| GUAAU | 2 |
| AUCCC | 1 |
| CCACCA | 1 |
| UUAAAA | 1 |
| GUAAUG | 1 |
| UUCAAGU | 1 |
| GUGACU | 1 |
| UUCGC | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- T-loop related (5)
- (4)
- T-loop with 1 bulged base (2)
- Other HL (1)
- T-loop with 1 bulged base (H) (1)
- Other HL (H) (1)
- Basepair signature
- cWW-F-F-F-F-F
- Heat map statistics
- Min 0.09 | Avg 0.48 | Max 0.99
Coloring options: