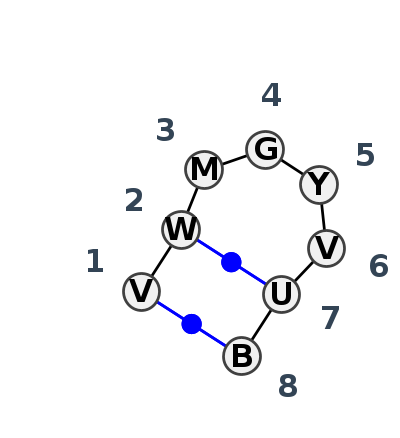
Motif HL_76195.1 Version HL_76195.1 of this group appears in releases 4.1 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 1-8 | 2-7 | 3-6 | 3-7 | 4-6 | 6-7 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_7A0S_009 | 7A0S | 0.6079 | 4 | X | LSU rRNA | A | 254 | A | 255 | C | 256 | G | 257 | C | 258 | G | 263 | U | 264 | U | 265 | ncWW | ncSH | ||||
| 2 | HL_5E6M_001 | 5E6M | 0.0000 | 3 | C | tRNA | G | 12 | U | 13 | A | 14 | G | 15 | U | 16 | A | 21 | U | 22 | C | 23 | cWW | cwW | cWS | |||
| 3 | HL_4QEI_001 | 4QEI | 0.1172 | 3 | C | tRNA | G | 12 | U | 13 | A | 14 | G | 15 | U | 16 | A | 21 | U | 22 | C | 23 | cWW | cwW | cWS | |||
| 4 | HL_2D6F_004 | 2D6F | 0.2338 | 4 | F | tRNA | G | 912 | U | 913 | A | 914 | G | 915 | U | 916 | A | 921 | U | 922 | C | 923 | cWW | ncwW | ncWS | |||
| 5 | HL_4WF9_068 | 4WF9 | 0.7796 | 4 | X | LSU rRNA | C | 280 | A | 281 | A | 282 | G | 283 | C | 284 | C | 288 | U | 289 | G | 291 | cWW | ncWW | ncWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GUAGUGGUAUC | 2 |
| AACGCUUGCGUU | 1 |
| GUAGUGGUAAUC | 1 |
| CAAGCUUGCUUG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UAGUGGUAU | 2 |
| ACGCUUGCGU | 1 |
| UAGUGGUAAU | 1 |
| AAGCUUGCUU | 1 |
Release history
| Release | 4.1 | 4.2 | 4.3 | 4.4 | 4.5 | 4.6 | 4.7 |
|---|---|---|---|---|---|---|---|
| Date | 2025-09-10 | 2025-10-08 | 2025-11-05 | 2025-12-03 | 2025-12-31 | 2026-01-28 | 2026-02-25 |
| Status | New id, 2 parents | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (5)
- Basepair signature
- cWW-cWW-F-F-F-F
- Heat map statistics
- Min 0.12 | Avg 0.45 | Max 0.82
Coloring options: