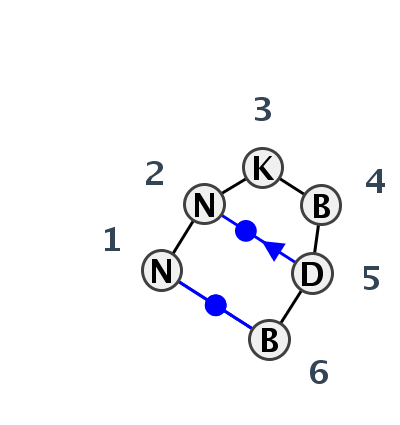
Motif HL_84503.1 Version HL_84503.1 of this group appears in releases 3.77 to 3.79
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 1-6 | 2-5 | 3-4 | 3-5 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_1N78_001 | 1N78 | 0.8924 | 4 | C | tRNA | U | 513 | A | 514 | G | 515 | U | 520 | A | 521 | G | 522 | cWW | cWS | ntSH | |
| 2 | HL_2DER_004 | 2DER | 0.6459 | 5 | D | tRNA | U | 13 | A | 14 | G | 15 | C | 20 | A | 21 | G | 22 | cWW | cWS | tSW | |
| 3 | HL_5HR6_001 | 5HR6 | 0.6364 | 5 | C | tRNA | U | 12 | A | 13 | G | 14 | C | 19 | A | 21 | G | 22 | cWW | cWS | tSW | |
| 4 | HL_5HR7_004 | 5HR7 | 0.6439 | 5 | D | tRNA | U | 13 | A | 14 | G | 15 | C | 20 | A | 21 | G | 22 | cWW | cWS | tSW | |
| 5 | HL_1QU2_002 | 1QU2 | 0.0000 | 3 | T | tRNA | C | 31 | C | 32 | U | 33 | U | 36 | A | 38 | G | 39 | cWW | ncWW | ntSW | |
| 6 | HL_3DIL_004 | 3DIL | 0.4966 | 0 | A | Lysine riboswitch | G | 149 | G | 150 | U | 151 | C | 152 | U | 153 | C | 154 | cWW | ncSW | ||
| 7 | HL_8S95_004 | 8S95 | 0.5276 | 2 | C | Enterovirus 5' cloverleaf cis-acting replication element | A | 91 | U | 92 | U | 93 | G | 96 | G | 97 | U | 98 | cWW | ncWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UAGAGGCCCAG | 3 |
| UAGCGGUUAG | 1 |
| CCUGAUAAG | 1 |
| GGUCUC | 1 |
| AUUGCGGU | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AGAGGCCCA | 3 |
| AGCGGUUA | 1 |
| CUGAUAA | 1 |
| GUCU | 1 |
| UUGCGG | 1 |
- Annotations
-
- (7)
- Basepair signature
- cWW-cWS-F-F
- Heat map statistics
- Min 0.07 | Avg 0.52 | Max 0.94
Coloring options: