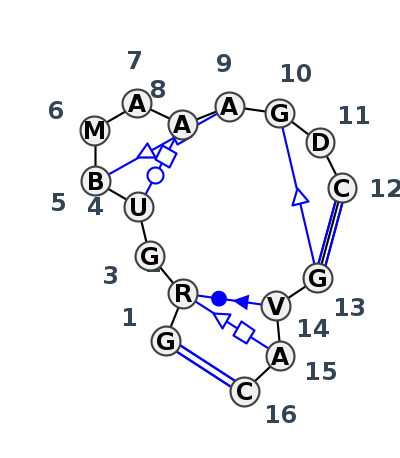
Motif IL_44923.2 Version IL_44923.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | break | 13 | 14 | 15 | 16 | 1-16 | 2-14 | 2-15 | 3-10 | 4-8 | 5-9 | 10-13 | 11-14 | 12-13 | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_8P9A_243 | 8P9A | 0.1282 | 2 | AR | LSU rRNA | G | 390 | A | 391 | G | 392 | U | 393 | G | 394 | A | 395 | A | 396 | A | 397 | A | 399 | G | 400 | U | 401 | C | 403 | * | G | 376 | A | 377 | A | 378 | C | 379 | cWW | cWS | tSH | cWB | tWH | tSH | tSS | cWW | |
| 2 | IL_8OI5_015 | 8OI5 | 0.1262 | 2 | 1 | LSU rRNA | G | 390 | A | 391 | G | 392 | U | 393 | G | 394 | A | 395 | A | 396 | A | 397 | A | 399 | G | 400 | U | 401 | C | 403 | * | G | 376 | A | 377 | A | 378 | C | 379 | cWW | ncWS | tSH | cWB | tWH | tSH | ntSS | cWW | |
| 3 | IL_9H3G_017 | 9H3G | 0.0855 | 2 | A | LSU rRNA | G | 387 | A | 388 | G | 389 | U | 390 | C | 391 | A | 392 | A | 393 | A | 394 | A | 396 | G | 397 | U | 398 | C | 400 | * | G | 373 | G | 374 | A | 375 | C | 376 | cWW | ncWS | tSH | cWB | tWH | ntSH | ntSS | cWW | |
| 4 | IL_8GLP_019 | 8GLP | 0.0000 | 2 | L5 | LSU rRNA | G | 401 | A | 402 | G | 403 | U | 404 | U | 405 | C | 406 | A | 407 | A | 408 | A | 410 | G | 411 | G | 412 | C | 414 | * | G | 387 | A | 388 | A | 389 | C | 390 | cWW | cWS | tSH | cWB | tWH | ntSH | ntSS | cWW | |
| 5 | IL_9AXU_014 | 9AXU | 0.1304 | 2 | 2 | LSU rRNA | G | 398 | A | 399 | G | 400 | U | 401 | U | 402 | A | 403 | A | 404 | A | 405 | A | 407 | G | 408 | U | 409 | C | 411 | * | G | 384 | A | 385 | A | 386 | C | 387 | cWW | cWS | tSH | cWB | tWH | ntSH | ntSS | cWW | |
| 6 | IL_9E6Q_018 | 9E6Q | 0.1615 | 1 | 1 | LSU rRNA | G | 526 | G | 527 | G | 528 | U | 529 | G | 530 | A | 531 | A | 532 | A | 533 | A | 534 | G | 535 | A | 536 | C | 538 | * | G | 512 | A | 513 | A | 514 | C | 515 | cWW | tSH | cWB | tWH | tSH | tSS | ntHH | cWW | |
| 7 | IL_4WF9_015 | 4WF9 | 0.1658 | 2 | X | LSU rRNA | G | 541 | A | 542 | G | 543 | U | 544 | G | 545 | A | 546 | A | 547 | A | 548 | A | 550 | G | 551 | A | 552 | C | 554 | * | G | 527 | C | 528 | A | 529 | C | 530 | cWW | cWS | tSH | cWB | tWH | ntSH | tSS | cWW | |
| 8 | IL_9DFE_012 | 9DFE | 0.1912 | 2 | 1A | LSU rRNA | G | 496 | A | 497 | G | 498 | U | 499 | G | 500 | A | 501 | A | 502 | A | 503 | A | 505 | G | 506 | A | 507 | C | 509 | * | G | 481 | A | 482 | A | 483 | C | 484 | cWW | cWS | tSH | cWB | tWH | tSH | tSS | ntHH | cWW |
| 9 | IL_8B0X_075 | 8B0X | 0.1885 | 2 | a | LSU rRNA | G | 496 | A | 497 | G | 498 | U | 499 | G | 500 | A | 501 | A | 502 | A | 503 | A | 505 | G | 506 | A | 507 | C | 509 | * | G | 481 | A | 482 | A | 483 | C | 484 | cWW | cWS | tSH | cWB | tWH | tSH | tSS | ntHH | cWW |
| 10 | IL_7A0S_009 | 7A0S | 0.2493 | 2 | X | LSU rRNA | G | 506 | A | 507 | G | 508 | U | 509 | G | 510 | A | 511 | A | 512 | A | 513 | A | 515 | G | 516 | A | 517 | C | 519 | * | G | 492 | A | 493 | A | 494 | C | 495 | cWW | cWS | tSH | cWB | tWH | ntSH | tSS | ntHH | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GAGUGAAAAAGUAC*GAAC | 2 |
| GAGUCAAAGAGUGC*GGAC | 1 |
| GAGUUCAAGAGGGC*GAAC | 1 |
| GAGUUAAAUAGUAC*GAAC | 1 |
| GGGUGAAAAGAGC*GAAC | 1 |
| GAGUGAAAUAGAAC*GCAC | 1 |
| GAGUGAAAUAGAGC*GAAC | 1 |
| GAGUGAAAAAGAAC*GAAC | 1 |
| GAGUGAAAGAGAAC*GAAC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| AGUGAAAAAGUA*AA | 2 |
| AGUCAAAGAGUG*GA | 1 |
| AGUUCAAGAGGG*AA | 1 |
| AGUUAAAUAGUA*AA | 1 |
| GGUGAAAAGAG*AA | 1 |
| AGUGAAAUAGAA*CA | 1 |
| AGUGAAAUAGAG*AA | 1 |
| AGUGAAAAAGAA*AA | 1 |
| AGUGAAAGAGAA*AA | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (10)
- Basepair signature
- cWW-cWS-tSH-L-tWH-cWW-tSS-tSH-L-R-L
- Heat map statistics
- Min 0.09 | Avg 0.17 | Max 0.35
Coloring options: