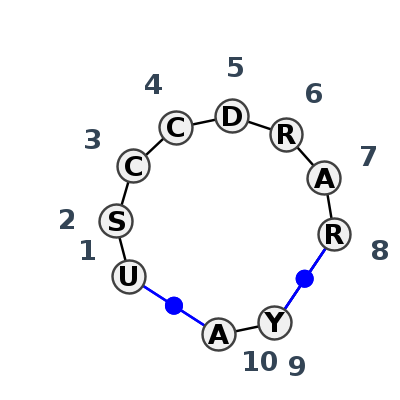
Motif IL_46868.1 Version IL_46868.1 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | break | 9 | 10 | 1-10 | 5-6 | 8-9 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IL_4WF9_037 | 4WF9 | 0.0766 | 0 | SSU/LSU pseudoknot | X | LSU rRNA | U | 1048 | C | 1049 | C | 1050 | C | 1051 | A | 1052 | A | 1053 | A | 1054 | A | 1055 | * | U | 1194 | A | 1195 | cWW | ncSH | cWW |
| 2 | IL_8B0X_098 | 8B0X | 0.0000 | 0 | a | LSU rRNA | U | 1006 | C | 1007 | C | 1008 | C | 1009 | A | 1010 | A | 1011 | A | 1012 | G | 1013 | * | C | 1152 | A | 1153 | cWW | ncSH | cWW | |
| 3 | IL_7A0S_034 | 7A0S | 0.1594 | 0 | SSU/LSU pseudoknot | X | LSU rRNA | U | 1015 | C | 1016 | C | 1017 | C | 1018 | U | 1019 | A | 1020 | A | 1021 | A | 1022 | * | U | 1161 | A | 1162 | cWW | ncSH | cWW |
| 4 | IL_9AXU_046 | 9AXU | 0.2224 | 0 | SSU/LSU pseudoknot (H) | 2 | LSU rRNA | U | 1204 | G | 1205 | C | 1206 | C | 1207 | G | 1208 | G | 1209 | A | 1210 | A | 1211 | * | U | 1356 | A | 1357 | cWW | chH | cWW |
| 5 | IL_8P9A_277 | 8P9A | 0.2489 | 0 | SSU/LSU pseudoknot | AR | LSU rRNA | U | 1173 | G | 1174 | C | 1175 | C | 1176 | G | 1177 | G | 1178 | A | 1179 | A | 1180 | * | U | 1325 | A | 1326 | cWW | nchH | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| UGCCGGAA*UA | 2 |
| UCCCAAAA*UA | 1 |
| UCCCAAAG*CA | 1 |
| UCCCUAAA*UA | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| CCCAAA* | 2 |
| GCCGGA* | 2 |
| CCCUAA* | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | New id, 2 parents |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- SSU/LSU pseudoknot (3)
- (1)
- SSU/LSU pseudoknot (H) (1)
- Basepair signature
- cWW-L-cWW-L-L-R-L-R
- Heat map statistics
- Min 0.05 | Avg 0.15 | Max 0.26
Coloring options: