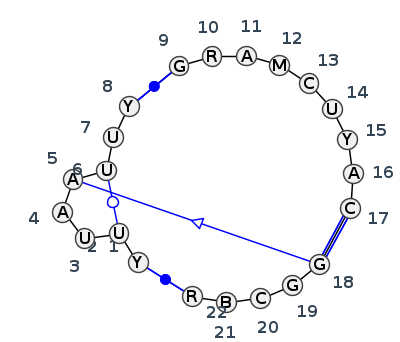
Motif J3_88451.2 Version J3_88451.2 of this group appears in releases 4.7 to 4.7
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | break | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | break | 18 | 19 | 20 | 21 | 22 | 1-22 | 2-6 | 5-17 | 5-18 | 8-9 | 14-20 | 15-19 | 15-20 | 17-18 | ||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | J3_8B0X_007 | 8B0X | 0.1293 | 3 | A | SSU rRNA | U | 955 | U | 956 | U | 957 | A | 958 | A | 959 | U | 960 | U | 961 | C | 962 | * | G | 973 | A | 974 | A | 977 | C | 979 | C | 980 | U | 981 | U | 982 | A | 983 | C | 984 | * | G | 1221 | G | 1222 | C | 1223 | U | 1224 | A | 1225 | cWW | tWWa | tSs | cWW | ncWW | ncWW | cWW | ||
| 2 | J3_4LFB_004 | 4LFB | 0.1244 | 3 | A | SSU rRNA | U | 955 | U | 956 | U | 957 | A | 958 | A | 959 | U | 960 | U | 961 | C | 962 | * | G | 973 | A | 974 | A | 977 | C | 979 | C | 980 | U | 981 | U | 982 | A | 983 | C | 984 | * | G | 1221 | G | 1222 | C | 1223 | G | 1224 | A | 1225 | cWW | tWWa | tSs | cWW | ncWW | cWW | |||
| 3 | J3_9H3G_037 | 9H3G | 0.0733 | 3 | h1 | SSU rRNA | C | 1181 | PSU | 1182 | U | 1183 | A | 1184 | A | 1185 | U | 1186 | U | 1187 | PSU | 1188 | * | G | 1199 | G | 1201 | A | 1203 | A | 1205 | C | 1206 | U | 1207 | PSU | 1208 | A | 1209 | C | 1210 | * | G | 1457 | G | 1458 | C | 1459 | C | 1460 | G | 1461 | cWW | tWWa | tSs | cWW | ncWW | ncWW | cWW | ||
| 4 | J3_8CRE_080 | 8CRE | 0.0000 | 3 | CM | SSU rRNA | C | 1165 | U | 1166 | U | 1167 | A | 1168 | A | 1169 | U | 1170 | U | 1171 | U | 1172 | * | G | 1183 | G | 1185 | A | 1187 | A | 1189 | C | 1190 | U | 1191 | C | 1192 | A | 1193 | C | 1194 | * | G | 1440 | G | 1441 | C | 1442 | C | 1443 | G | 1444 | cWW | tWWa | tSs | cWW | cWW | ||||
| 5 | J3_8P9A_078 | 8P9A | 0.0772 | 3 | sR | SSU rRNA | C | 1180 | U | 1181 | U | 1182 | A | 1183 | A | 1184 | U | 1185 | U | 1186 | U | 1187 | * | G | 1198 | G | 1200 | A | 1202 | A | 1204 | C | 1205 | U | 1206 | C | 1207 | A | 1208 | C | 1209 | * | G | 1454 | G | 1455 | C | 1456 | C | 1457 | G | 1458 | cWW | tWWa | ncSs | tSs | cWW | cWW | |||
| 6 | J3_8GLP_046 | 8GLP | 0.1947 | 3 | S2 | SSU rRNA | C | 1237 | PSU | 1238 | PSU | 1239 | A | 1240 | A | 1241 | U | 1242 | PSU | 1243 | PSU | 1244 | * | G | 1255 | G | 1257 | A | 1259 | C | 1261 | C | 1262 | U | 1263 | C | 1264 | A | 1265 | C | 1266 | * | G | 1516 | G | 1517 | C | 1518 | U | 1519 | G | 1520 | cWW | tWWa | cSs | tSs | cWW | ncWW | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| CUUAAUUU*GGGGAAACUCAC*GGCCG | 2 |
| UUUAAUUC*GAAGAACCUUAC*GGCUA | 1 |
| UUUAAUUC*GAAGAACCUUAC*GGCGA | 1 |
| C(PSU)UAAUUPSU*GGGGAAACU(PSU)AC*GGCCG | 1 |
| C(PSU)(PSU)AAU(PSU)PSU*GGGAAACCUCAC*GGCUG | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UUAAUU*GGGAAACUCA*GCC | 2 |
| UUAAUU*AAGAACCUUA*GCU | 1 |
| UUAAUU*AAGAACCUUA*GCG | 1 |
| (PSU)UAAUU*GGGAAACU(PSU)A*GCC | 1 |
| (PSU)(PSU)AAU(PSU)*GGAAACCUCA*GCU | 1 |
Release history
| Release | 4.7 |
|---|---|
| Date | 2026-02-25 |
| Status | Updated, 1 parent |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- (6)
- Basepair signature
- cWW-tWW-F-F-F-F-F-tSS-cWW-F-F-cWW-F-F-F-F-F-F
- Heat map statistics
- Min 0.05 | Avg 0.12 | Max 0.26
Coloring options: