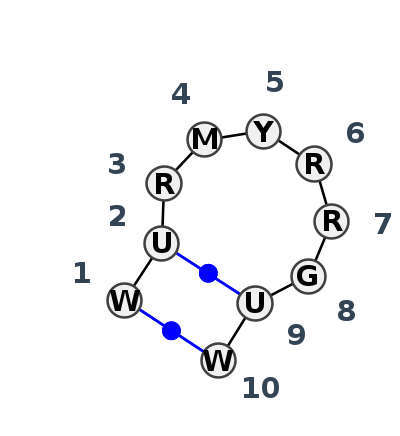
Motif HL_35704.1 Version HL_35704.1 of this group appears in releases 4.1 to 4.6
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 1-10 | 2-9 | 4-7 | 5-7 | 8-9 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_4ATO_001 | 4ATO | 0.5870 | 1 | G | ToxI antitoxin | U | 15 | U | 16 | A | 17 | C | 18 | C | 19 | G | 20 | A | 22 | G | 23 | U | 24 | A | 25 | cWW | cwW | cSH | ||
| 2 | HL_9UW0_001 | 9UW0 | 0.1033 | 1 | X | RNA (65-MER) | A | 19 | U | 20 | G | 21 | A | 22 | U | 23 | A | 24 | G | 26 | G | 27 | U | 28 | U | 29 | cWW | cwW | ncSs | ||
| 3 | HL_5SWD_004 | 5SWD | 0.0794 | 1 | B | Purine riboswitch | A | 30 | U | 31 | G | 32 | A | 33 | U | 34 | A | 35 | G | 37 | G | 38 | U | 39 | U | 40 | cWW | cwW | ncSs | ||
| 4 | HL_3LA5_001 | 3LA5 | 0.0000 | 1 | A | Purine riboswitch | A | 30 | U | 31 | G | 32 | A | 33 | U | 34 | A | 35 | G | 37 | G | 38 | U | 39 | U | 40 | cWW | cwW | ncSs | ||
| 5 | HL_4LX6_001 | 4LX6 | 0.0835 | 1 | A | Purine riboswitch | A | 30 | U | 31 | G | 32 | A | 33 | U | 34 | A | 35 | G | 37 | G | 38 | U | 39 | U | 40 | cWW | cwW | ncSs | ||
| 6 | HL_4XNR_001 | 4XNR | 0.0958 | 1 | X | Purine riboswitch | A | 30 | U | 31 | G | 32 | A | 33 | U | 34 | A | 35 | G | 37 | G | 38 | U | 39 | U | 40 | cWW | cwW | ncSs | ||
| 7 | HL_1Y26_001 | 1Y26 | 0.1152 | 1 | X | Purine riboswitch | A | 30 | U | 31 | G | 32 | A | 33 | U | 34 | A | 35 | G | 37 | G | 38 | U | 39 | U | 40 | cWW | cwW | ncsS | ncSs | |
| 8 | HL_8KEB_001 | 8KEB | 0.0930 | 1 | A | RNA (72-MER) | A | 18 | U | 19 | G | 20 | A | 21 | U | 22 | A | 23 | G | 25 | G | 26 | U | 27 | U | 28 | cWW | cwW | ncsS | ||
| 9 | HL_9LKV_001 | 9LKV | 0.0850 | 1 | X | RNA (73-MER) | A | 19 | U | 20 | G | 21 | A | 22 | U | 23 | A | 24 | G | 26 | G | 27 | U | 28 | U | 29 | cWW | cwW | ncSs | ||
| 10 | HL_3IVN_001 | 3IVN | 0.1713 | 1 | A | Purine riboswitch | A | 21 | U | 22 | G | 23 | A | 24 | U | 25 | A | 26 | G | 28 | G | 29 | U | 30 | U | 31 | cWW | cwW | ncSs | ||
| 11 | HL_7KVT_001 | 7KVT | 0.2311 | 1 | B | Squash RNA aptamer bound to DFHBI-1T with iridium (III) ions | A | 23 | U | 24 | A | 25 | A | 26 | U | 27 | A | 28 | G | 30 | G | 31 | U | 32 | U | 33 | cWW | cwW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| AUGAUAUGGUU | 9 |
| UUACCGUAGUA | 1 |
| AUAAUAUGGUU | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| UGAUAUGGU | 9 |
| UACCGUAGU | 1 |
| UAAUAUGGU | 1 |
- Annotations
-
- (11)
- Basepair signature
- cWW-cWW-F-F-F-F-F-F
- Heat map statistics
- Min 0.06 | Avg 0.21 | Max 0.63
Coloring options: