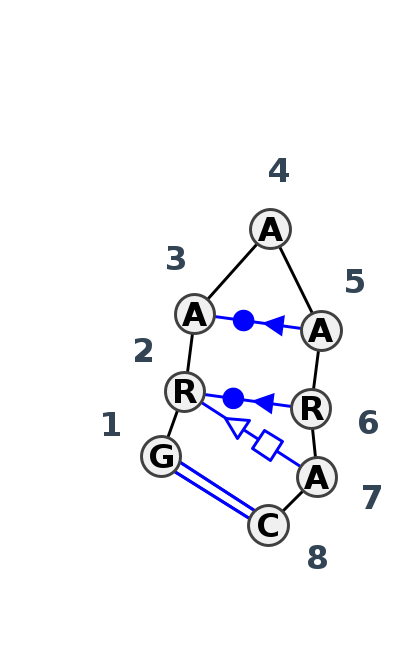
Motif HL_68767.1 Version HL_68767.1 of this group appears in releases 3.89 to 3.99
| #S | Loop id | PDB | Disc | #Non-core | Annotation | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 1-8 | 2-6 | 2-7 | 3-5 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_2BTE_001 | 2BTE | 0.1053 | 5 | tRNA D-loop | B | tRNA | G | 12 | G | 13 | A | 14 | A | 15 | A | 20|||A | G | 21 | A | 22 | C | 23 | cWW | cWS | tSH | cWS |
| 2 | HL_3ZGZ_001 | 3ZGZ | 0.0000 | 5 | tRNA D-loop | B | tRNA | G | 12 | G | 13 | A | 14 | A | 15 | A | 20|||A | G | 21 | A | 22 | C | 23 | cWW | cWS | tSH | cWS |
| 3 | HL_5AH5_005 | 5AH5 | 0.1111 | 5 | tRNA D-loop | D | tRNA | G | 12 | A | 13 | A | 14 | A | 15 | A | 20|||A | A | 21 | A | 22 | C | 23 | cWW | cWS | tSH | cWS |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| GGAAUGGGUAGAC | 1 |
| GGAAUCGGUAGAC | 1 |
| GAAAUCGGUAAAC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| GAAUGGGUAGA | 1 |
| GAAUCGGUAGA | 1 |
| AAAUCGGUAAA | 1 |
Release history
| Release | 3.89 | 3.90 | 3.91 | 3.92 | 3.93 | 3.94 | 3.95 | 3.96 | 3.97 | 3.98 | 3.99 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Date | 2024-10-09 | 2024-11-06 | 2024-12-04 | 2025-01-01 | 2025-01-29 | 2025-02-26 | 2025-03-26 | 2025-04-23 | 2025-05-21 | 2025-06-18 | 2025-07-16 |
| Status | New id, 1 parent | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match | Exact match |
Parent motifs
This motif has no parent motifs.
Children motifs
This motif has no children motifs.- Annotations
-
- tRNA D-loop (3)
- Basepair signature
- cWW-cWS-tSH-cWS-F
- Heat map statistics
- Min 0.11 | Avg 0.08 | Max 0.13
Coloring options: