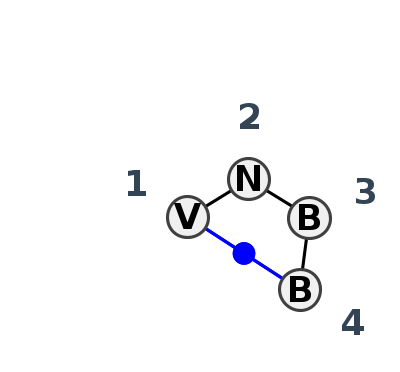
Motif HL_97696.1 Version HL_97696.1 of this group appears in releases 4.1 to 4.6
| #S | Loop id | PDB | Disc | #Non-core | Chain(s) | Standardized name for chain | 1 | 2 | 3 | 4 | 1-4 | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HL_9E6Q_071 | 9E6Q | 0.6199 | 0 | 1 | LSU rRNA | G | 1678 | U | 1679 | U | 1680 | U | 1681 | ncWW |
| 2 | HL_8OI5_012 | 8OI5 | 0.5715 | 4 | 1 | LSU rRNA | G | 539 | C | 540 | U | 541 | C | 546 | cWW |
| 3 | HL_3SKI_003 | 3SKI | 0.0000 | 2 | A | 2dG-I riboswitch aptamer | A | 67 | C | 68 | C | 69 | U | 72 | cWW |
| 4 | HL_3SKL_007 | 3SKL | 0.1729 | 2 | B | 2dG-I riboswitch aptamer | A | 66 | C | 67 | C | 68 | U | 71 | ncWW |
| 5 | HL_2Y9H_004 | 2Y9H | 0.2309 | 2 | H | RNA (19-mer) | C | 11 | G | 12 | C | 13 | G | 16 | cWW |
| 6 | HL_4KJI_001 | 4KJI | 0.4607 | 4 | C | RsmZ-2 | G | 5 | A | 6 | G | 8 | C | 12 | cWW |
3D structures
Complete motif including flanking bases
| Sequence | Counts |
|---|---|
| ACCUAU | 2 |
| GUUU | 1 |
| GCUUCUGC | 1 |
| CGCGUG | 1 |
| GAAGGAUC | 1 |
Non-Watson-Crick part of the motif
| Sequence | Counts |
|---|---|
| CCUA | 2 |
| UU | 1 |
| CUUCUG | 1 |
| GCGU | 1 |
| AAGGAU | 1 |
Coloring options: